Summary: Researchers in China successfully used CRISPR gene editing to produce five monkey clones from the fibroblasts of a donor monkey with disease phenotypes.
Source: Science China Press
The first cohort of five gene-edited monkey clones made from fibroblasts of a monkey with disease phenotypes were born recently at the Institute of Neuroscience (ION) of Chinese Academy of Sciences (CAS) in Shanghai.
The expression of BMAL1, a core circadian regulatory transcription factor, was knockout in the donor monkey using CRISPR/Cas9-mediated gene editing at the embryo stage, and the fibroblasts of the donor monkey were used to clone five monkeys using the method of somatic cell nuclear transfer, the same method that generated Zhong Zhong and Hua Hua, the first two cloned monkeys, last year.
This major advance, reported on-line in two articles in the journal National Science Review on January 24, demonstrates that a population of customized gene-edited macaque monkeys with uniform genetic background will soon be available for biomedical research.
The first article describes the generation of gene-edited donor monkeys, using CRISPR-Cas9 method to edit the BMAL1 gene of in vitro fertilized monkey embryos. These monkeys exhibited a wide-range of circadian disorder phenotypes, including reduced sleep time, elevated night-time locomotive activities, dampened circadian cycling of blood hormones, increased anxiety and depression, as well as schizophrenia-like behaviors.
“Disorder of circadian rhythm could lead to many human diseases, including sleep disorders, diabetic mellitus, cancer, and neurodegenerative diseases, our BMAL1-knock out monkeys thus could be used to study the disease pathogenesis as well as therapeutic treatments” says Hung-Chun Chang, senior author and investigator of the Chinese Academy of Sciences Institute of Neuroscience.
The second article describes the cloning of macaque monkeys from the fibroblast of a BMAL1-knockout monkey, using the method of somatic cell nuclear transfer.
In this method, the researchers removed the nucleus from a monkey oocyte (egg cell) and replaced it with another nucleus from a fibroblast, a differentiated somatic (body) cell. This reconstructed egg then developed into an embryo that carries the genes of the replacement nucleus. The embryo was then transfer to the womb of a surrogate female monkey that later gave birth to the cloned monkey.
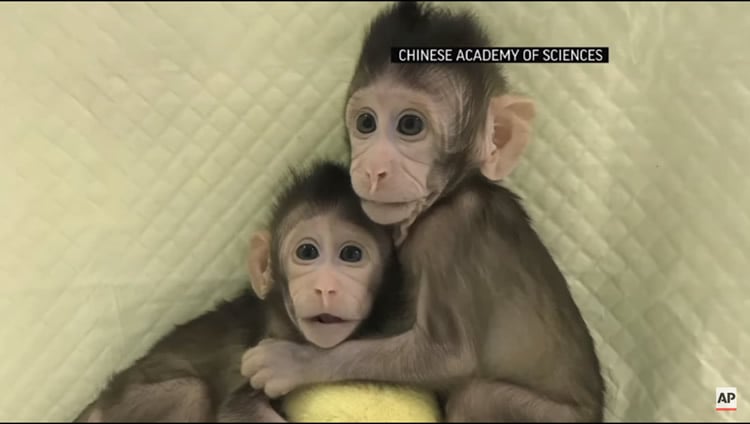

In the previous work, Zhong Zhong and Hua Hua were generated by using fibroblasts from an aborted fetus. The present work succeeded in using fibroblasts from a young adult gene-edited donor monkey with disease phenotypes.
“Our approach is to perform gene-editing in fertilized embryos to first generate a group of gene-edited monkeys, and then select one monkey that exhibits correct gene editing and most severe disease phenotypes as the donor monkey for cloning by somatic cell nuclear transfer” says Qiang Sun, senior author of the paper and Director of ION’s Nonhuman Primate Research Facility.
“We believe that this approach of cloning gene-edited monkeys could be used to generate a variety of monkey models for gene-based diseases, including many brain diseases, as well as immune and metabolic disorders and cancer”.
The researchers plan to continue improving the technique in order to increase the efficiency of cloning. The group is expecting more macaque clones carrying disease-causing gene mutations to be generated in the coming years.
Video credited to Associated Press via YouTube
The Institute of Neuroscience, CAS is following strict international guidelines for animal research.
“This work required coordinated efforts of many laboratories, and serves as a clear example of the efficient team work that is highly emphasized by CAS” says Mu-ming Poo, A co-author on both studies, who directs the Institute of Neuroscience and helps to supervise the project. “This line of research will help to reduce the amount of macaque monkeys currently used in biomedical research around the world”.
“Without the interference of genetic background, a much smaller number of cloned monkeys carrying disease phenotypes may be sufficient for pre-clinical tests of the efficacy of therapeutics,” Poo says.
Funding: This work was supported by grants from Chinese Academy of Sciences, Shanghai Municipal Government Bureau of Science and Technology, and Ministry of Science and Technology of China.
Source:
Science China Press
Media Contacts:
WANG Zuoren – Science China Press
Image Source:
The image is adapted from the Associated Press video.
Original Research: Open access
“BMAL1 knockout macaque monkeys display reduced sleep and psychiatric disorders”
Peiyuan Qiu, Jian Jiang, Zhen Liu, Yijun Cai, Tao Huang, Yan Wang Qiming Liu, Yanhong Nie, Fang Liu, Jiumu Cheng, Qing Li, Yun-Chi Tang, Mu-ming Poo, Qiang Sun, Hung-Chun Chang. National Science Review, Volume 6, Issue 1, January 2019, Pages 87–100, doi:10.1093/nsr/nwz002
Open access
“Cloning of a gene-edited macaque monkey by somatic cell nuclear transfer”
Zhen Liu, Yijun Cai, Zhaodi Liao, Yuting Xu, Yan Wang, Zhanyang Wang, Xiaoyu Jiang, Yuzhuo Li, Yong Lu, Yanhong Nie, Xiaotong Zhang, Chunyang Li, Xinyan Bian, Mu-ming Poo, Hung-Chun Chang, Qiang Sun.
National Science Review, Volume 6, Issue 1, January 2019, Pages 101–108, doi:10.1093/nsr/nwz003
Abstract
BMAL1 knockout macaque monkeys display reduced sleep and psychiatric disorders
Circadian disruption is a risk factor for metabolic, psychiatric and age-related disorders, and non-human primate models could help to develop therapeutic treatments. Here, we report the generation of BMAL1 knockout cynomolgus monkeys for circadian-related disorders by CRISPR/Cas9 editing of monkey embryos. These monkeys showed higher nocturnal locomotion and reduced sleep, which was further exacerbated by a constant light regimen. Physiological circadian disruption was reflected by the markedly dampened and arrhythmic blood hormonal levels. Furthermore, BMAL1-deficient monkeys exhibited anxiety and depression, consistent with their stably elevated blood cortisol, and defective sensory processing in auditory oddball tests found in schizophrenia patients. Ablation of BMAL1 up-regulated transcriptional programs toward inflammatory and stress responses, with transcription networks associated with human sleep deprivation, major depressive disorders, and aging. Thus, BMAL1 knockout monkeys are potentially useful for studying the physiological consequences of circadian disturbance, and for developing therapies for circadian and psychiatric disorders.
Abstract
Cloning of a gene-edited macaque monkey by somatic cell nuclear transfer
Cloning of macaque monkeys by somatic cell nucleus transfer (SCNT) allows the generation of monkeys with uniform genetic backgrounds that are useful for the development of non-human primate models of human diseases. Here, we report the feasibility of this approach by SCNT of fibroblasts from a macaque monkey (Macaca fascicularis), in which a core circadian transcription factor BMAL1 was knocked out by clustered regularly interspaced short palindromic repeat/Cas9 gene editing (see accompanying paper). Out of 325 SCNT embryos transferred into 65 surrogate monkeys, we cloned five macaque monkeys with BMAL1 mutations in both alleles without mosaicism, with nuclear genes identical to that of the fibroblast donor monkey. Further peripheral blood mRNA analysis confirmed the complete absence of the wild-type BMAL1 transcript. This study demonstrates that the SCNT approach could be used to generate cloned monkeys from fibroblasts of a young adult monkey and paves the way for the development of macaque monkey disease models with uniform genetic backgrounds.