Key molecular pathways that ultimately lead to late-onset Alzheimer’s disease, the most common form of the disorder, have been identified by researchers at Columbia University Medical Center (CUMC). The study, which used a combination of systems biology and cell biology tools, presents a new approach to Alzheimer’s disease research and highlights several new potential drug targets. The paper was published today in the journal Nature.
Much of what is known about Alzheimer’s comes from laboratory studies of rare, early-onset, familial (inherited) forms of the disease. “Such studies have provided important clues as to the underlying disease process, but it’s unclear how these rare familial forms of Alzheimer’s relate to the common form of the disease,” said study leader Asa Abeliovich, MD, PhD, associate professor of pathology and cell biology and of neurology in the Taub Institute for Research on Alzheimer’s Disease and the Aging Brain at CUMC. “Most important, dozens of drugs that ‘work’ in mouse models of familial disease have ultimately failed when tested in patients with late-onset Alzheimer’s. This has driven us, and other laboratories, to pursue mechanisms of the common form of the disease.”

Non-familial Alzheimer’s is complex; it is thought to be caused by a combination of genetic and environmental risk factors, each having a modest effect individually. Using so-called genome-wide association studies (GWAS), prior reports have identified a handful of common genetic variants that increase the likelihood of Alzheimer’s. A key goal has been to understand how such common genetic variants function to impact the likelihood of Alzheimer’s.
In the current study, the CUMC researchers identified key molecular pathways that link such genetic risk factors to Alzheimer’s disease. The work combined cell biology studies with systems biology tools, which are based on computational analysis of the complex network of changes in the expression of genes in the at-risk human brain.
More specifically, the researchers first focused on the single most significant genetic factor that puts people at high risk for Alzheimer’s, called APOE4 (found in about a third of all individuals). People with one copy of this genetic variant have a three-fold increased risk of developing late-onset Alzheimer’s, while those with two copies have a ten-fold increased risk. “In this study,” said Dr. Abeliovich, “we initially asked: If we look at autopsy brain tissue from individuals at high risk for Alzheimer’s, is there a consistent pattern?”
Surprisingly, even in the absence of Alzheimer’s disease, brain tissue from individuals at high risk (who carried APOE4 in their genes) harbored certain changes reminiscent of those seen in full-blown Alzheimer’s disease,” said Dr. Abeliovich. “We therefore focused on trying to understand these changes, which seem to put people at risk. The brain changes we considered were based on ‘transcriptomics’—a broad molecular survey of the expression levels of the thousands of genes expressed in brain.”
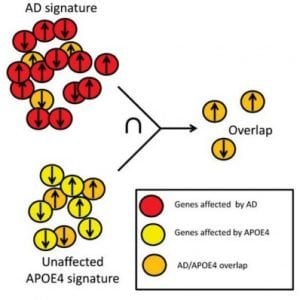
Using the network analysis tools mentioned above, the researchers then identified a dozen candidate “master regulator” factors that link APOE4 to the cascade of destructive events that culminates in Alzheimer’s dementia. Subsequent cell biology studies revealed that a number of these master regulators are involved in the processing and trafficking of amyloid precursor protein (APP) within brain neurons. APP gives rise to amyloid beta, the protein that accumulates in the brain cells of patients with Alzheimer’s. In sum, the work ultimately connected the dots between a common genetic factor that puts individuals at high risk for Alzheimer’s, APOE4, and the disease pathology.
Among the candidate “master regulators” identified, the team further analyzed two genes, SV2A and RFN219. “We were particularly interested in SV2A, as it is the target of a commonly used anti-epileptic drug, levetiracetam. This suggested a therapeutic strategy. But more research is needed before we can develop clinical trials of levetiracetam for patients with signs of late-onset Alzheimer’s disease.”
The researchers evaluated the role of SV2A, using human-induced neurons that carry the APOE4 genetic variant. (The neurons were generated by directed conversion of skin fibroblasts from individuals at high risk for Alzheimer’s, using a technology developed in the Abeliovich laboratory.) Treating neurons that harbor the APOE4 at-risk genetic variant with levetiracetam (which inhibits SV2A) led to reduced production of amyloid beta. The study also showed that RFN219 appears to play a role in APP-processing in cells with the APOE4 variant.

Notes about this Alzheimer’s disease research
The paper is titled, “Integrative genomics identifies APOE ɛ4 effectors in Alzheimer’s disease.” The other contributors are all members of CUMC’s Taub Institute for Research on Alzheimer’s Disease and the Aging Brain: Herve Rhinn, Ryousuke Fujita, Liang Qiang, Rong Chen, and Joseph H. Lee. Dr. Lee is also a member of CUMC’s Gertrude H. Sergievsky Center.
The authors declare no financial or other conflicts of interest. Subsequent to the acceptance of this paper, Dr. Abeliovich received a grant from UCB, Inc. for an unrelated study.
The study was supported by grants from the National Institutes of Health (R01AG042317 and RO1NS064433).
Contact: Press Office – Columbia University Medical Center
Source: Columbia University Medical Center
Image Source: All images are credited to Dr. Asa Abeliovich, CUMC and are adapted from the press release.
Original Research: Abstract for “Integrative genomics identifies APOE ε4 effectors in Alzheimer’s disease” by Herve Rhinn, Ryousuke Fujita, Liang Qiang, Rong Cheng, Joseph H. Lee and Asa Abeliovich in Nature. Published online July 24 2013 doi:10.1038/nature12415