Changes in the epigenome, including chemical modifications of DNA, can act as an extra layer of information in the genome, and are thought to play a role in learning and memory, as well as in age-related cognitive decline.
The results of a new study by scientists at the Salk Institute for Biological Studies show that the landscape of DNA methylation, a particular type of epigenomic modification, is highly dynamic in brain cells during the transition from birth to adulthood, helping to understand how information in the genomes of cells in the brain is controlled from fetal development to adulthood. The brain is much more complex than all other organs in the body and this discovery opens the door to a deeper understanding of how the intricate patterns of connectivity in the brain are formed.
“These results extend our knowledge of the unique role of DNA methylation in brain development and function,” says senior author Joseph R. Ecker, professor and director of Salk’s Genomic Analysis Laboratory and holder of the Salk International Council Chair in Genetics. “They offer a new framework for testing the role of the epigenome in healthy function and in pathological disruptions of neural circuits.”
A healthy brain is the product of a long process of development. The front-most part of our brain, called the frontal cortex, plays a key role in our ability to think, decide and act. The brain accomplishes all of this through the interaction of special cells such as neurons and glia. We know that these cells have distinct functions, but what gives these cells their individual identities? The answer lies in how each cell expresses the information contained in its DNA. Epigenomic modifications, such as DNA methylation, can control which genes are turned on or off without changing letters of the DNA alphabet (A-T-C-G), and thus help distinguish different cell types.
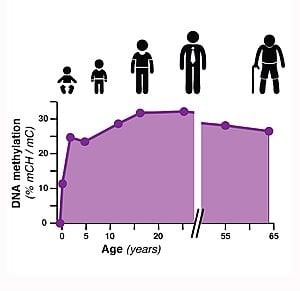
In this new study, published July 4, 2013, in Science, the scientists found that the patterns of DNA methylation undergo widespread reconfiguration in the frontal cortex of mouse and human brains during a time of development when synapses, or connections between nerve cells, are growing rapidly. The researchers identified the exact sites of DNA methylation throughout the genome in brains from infants through adults. They found that one form of DNA methylation is present in neurons and glia from birth. Strikingly, a second form of “non-CG” DNA methylation that is almost exclusive to neurons accumulates as the brain matures, becoming the dominant form of methylation in the genome of human neurons. These results help us to understand how the intricate DNA landscape of brain cells develops during the key stages of childhood.
The genetic code in DNA is made up of four chemical bases: adenine (A), guanine (G), cytosine (C), and thymine (T). DNA methylation typically occurs at so-called CpG sites, where C (cytosine) sits next to G (guanine) in the DNA alphabet. About 80 to 90 percent of CpG sites are methylated in human DNA. Salk researchers previously discovered that in human embryonic stem cells and induced pluripotent stem cells, a type of artificially derived stem cell, DNA methylation can also occur when G does not follow C, hence “non-CG methylation.” Originally, they thought that this type of methylation disappeared when stem cells differentiate into specific tissue-types such as lung or fat cells. The current study finds this is not the case in the brain, where non-CG methylation appears after cells differentiate, usually during childhood and adolescence when the brain is maturing.
By sequencing the genomes of mouse and human brain tissue as well as neurons and glia (from the frontal cortex of the brain) during early postnatal, juvenile, adolescent and adult stages, the Salk team found that non-CG methylation accumulates in neurons through early childhood and adolescence, and becomes the dominant form of DNA methylation in mature human neurons. “This shows that the period during which the neural circuits of the brain mature is accompanied by a parallel process of large-scale reconfiguration of the neural epigenome,” says Ecker, who is a Howard Hughes Medical Institute and Gordon and Betty Moore Foundation investigator.
The study provides the first comprehensive maps of how DNA methylation patterns change in the mouse and human brain during development, forming a critical foundation to now explore whether changes in methylation patterns may be linked to human diseases, including psychiatric disorders. Recent studies have demonstrated a possible role for DNA methylation in schizophrenia, depression, suicide and bipolar disorder. “Our work will let us begin to ask more detailed questions about how changes in the epigenome sculpt the complex identities of brain cells through life,” says co-first author Eran Mukamel, from Salk’s Computational Neurobiology Laboratory.
“The human brain has been called the most complex system that we know of in the universe,” says Ryan Lister, co-corresponding author on the new paper, previously a postdoctoral fellow in Ecker’s laboratory at Salk and now a group leader at The University of Western Australia. “So perhaps we shouldn’t be so surprised that this complexity extends to the level of the brain epigenome. These unique features of DNA methylation that emerge during critical phases of brain development suggest the presence of previously unrecognized regulatory processes that may be critically involved in normal brain function and brain disorders.”
At present, there is consensus among neuroscientists that many mental disorders have a neurodevelopmental origin and arise from an interaction between genetic predisposition and environmental influences (for example, early-life stress or drug abuse), the outcome of which is altered activity of brain networks. The building and shaping of these brain networks requires a long maturation process in which central nervous system cell types (neurons and glia) need to fine-tune the way they express their genetic code.
“DNA methylation fulfills this role,” says study co-author Terrence J. Sejnowski, a Howard Hughes Medical Institute Investigator, holder of the Francis Crick Chair and head of Salk’s Computational Neurobiology Laboratory. “We found that patterns of methylation are dynamic during brain development, in particular for non-CG methylation during early childhood and adolescence, which changes the way that we think about normal brain function and dysfunction.” By disrupting the transcriptional expression of neurons, adds co-corresponding author M. Margarita Behrens, a staff scientist in the Computational Neurobiology Laboratory, “the alterations of these methylation patterns will change the way in which networks are formed, which could, in turn, lead to the appearance of mental disorders later in life.”
Notes about this epigenomics and brain development research
Other researchers on the study were Joseph R. Nery, Mark Urich, Clare A. Puddifoot, Nicholas D. Johnson, Jacinto Lucero and Matthew D. Schultz from the Salk Institute; Julian Tonti-Filippini from the University of Western Australia; Yun Huang and Anjana Rao of the La Jolla Institute for Allergy and Immunology; Miao Yu and Chuan He of the University of Chicago; Andrew J. Dwork and Fatemeh G. Haghighi of Columbia University; Holger Heyn and Manel Esteller of Bellvigte Biomedical Research Institute; and Shijun Hu and Joseph C. Wu of the Stanford University School of Medicine.
The work was supported by the National Institute of Mental Health, the Howard Hughes Medical Institute, the Gordon and Betty Moore Foundation, the California Institute for Regenerative Medicine, the Leukemia and Lymphoma Society, the Australian Research Council, and the Center for Theoretical Biological Physics at the University of California, San Diego.
Contact: Kat Kearney – Salk Institute
Source: Salk Institute press release
Image Source: The epigenomics graph image is credited to Eran Mukamel and is adapted from the Salk Institute press release.
Original Research: Abstract for “Global Epigenomic Reconfiguration During Mammalian Brain Development” by Ryan Lister, Eran A. Mukamel, Joseph R. Nery, Mark Urich, Clare A. Puddifoot, Nicholas D. Johnson, Jacinta Lucero, Yun Huang, Andrew J. Dwork, Matthew D. Schultz, Miao Yu, Julian Tonti-Filippini, Holger Heyn, Shijun Hu, Joseph C. Wu, Anjana Rao, Manel Esteller, Chuan He, Fatemeh G. Haghighi, Terrence J. Sejnowski, M. Margarita Behrens, and Joseph R. Ecker in Science. Published online July 4 2013 DOI:10.1126/science.1237905