Summary: Researchers have identified causal links between microbiota gene content and metabolic activity which impacts personal differences in drug metabolism. The findings help explain why some people don’t respond to, or have adverse side effects from taking medications, while others don’t.
Source: Yale
Yale researchers identified human gut microbes that metabolize over 150 therapeutic drugs, a finding that highlights the role bacteria play in determining how well individuals respond to medications, they report June 3 in the journal Nature.
Scientists in the lab of senior author Andrew Goodman of Yale’s Microbial Sciences Institute and the Department of Microbial Pathogenesis also pinpointed the microbial genes responsible for many of these drug-metabolizing activities.
“It is possible that we can use genes or species of bacteria to predict the capacity of an individual’s gut flora to metabolize a certain drug,” said Maria Zimmermann-Kogadeeva, a postdoctoral fellow in Goodman’s lab and co-lead author of the study. “The work is a first step in identifying biomarkers that could help doctors prescribe the drugs that are the safest and most effective for individual patients.”
The metabolism of drugs was once thought to be exclusively the role of human organs such as the liver, which carry out the important task of turning chemical compounds into water-soluble molecules to help their elimination from the body. This process is crucial to determining the effectiveness and safety of clinical drugs: The drugs need to stay in the body long enough to have an effect but not long enough to accumulate and potentially become toxic.
Drug metabolism studies typically do not assess the contribution of the microbiome.
“However, anecdotal examples of microbiome-metabolized drugs have emerged over past decades,” Goodman said.
To map the connections between gut microbes and medical drugs, the researchers investigated whether and how each of 271 drugs are chemically modified by each of 76 different kinds of bacteria from the human gut. Almost two-thirds of these drugs were metabolized by at least one of the bacterial specimens tested. The researchers then constructed genetic libraries of the selected drug-metabolizing gut bacteria which allowed them to systematically identify many of the genes responsible for the chemical transformation of the drugs.
They found that the number of these genes varies widely across the gut microbial communities of healthy human volunteers. In some cases, these differences explain why some individuals’ gut microbiomes metabolize drugs rapidly, while others modify the same drugs slowly or not at all.
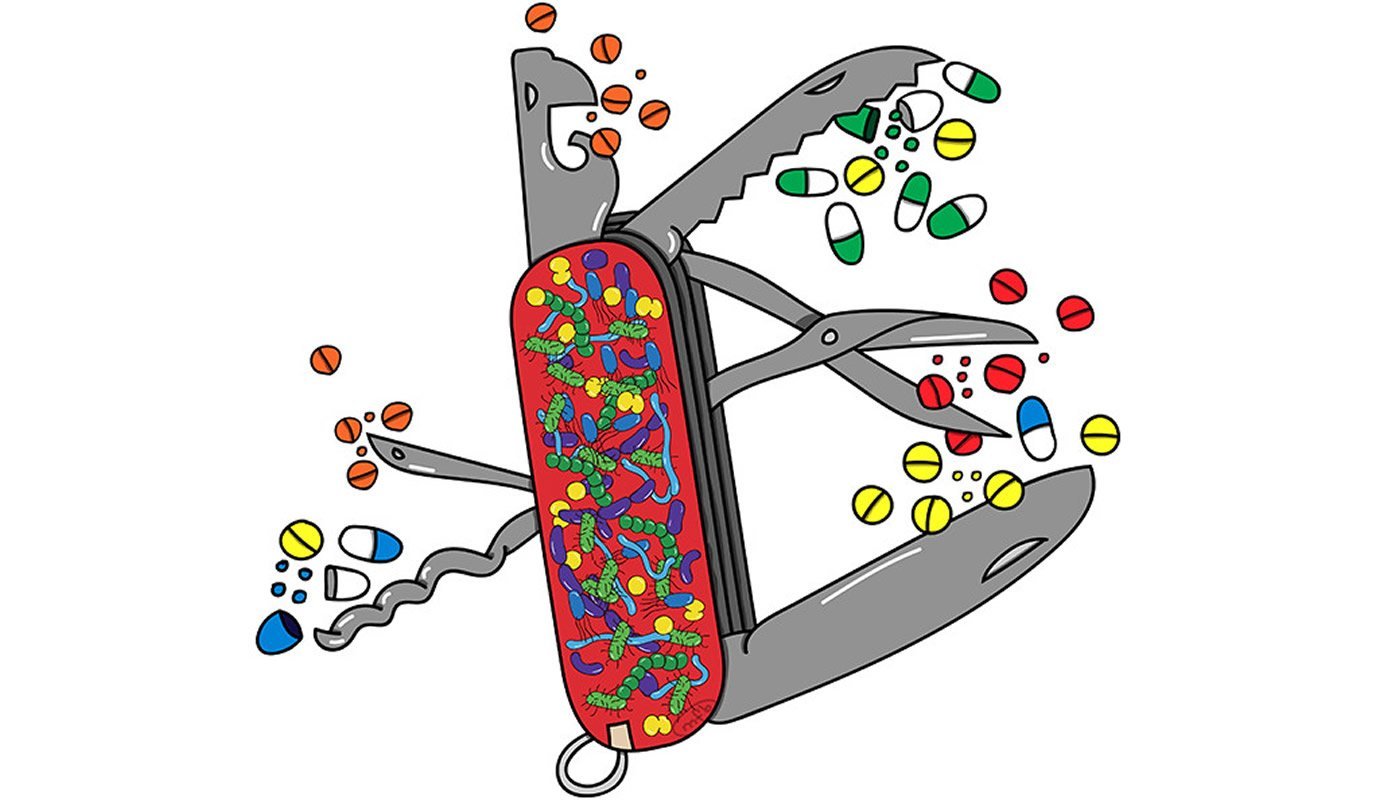
“We hope this study provides a useful first step in understanding the microbiome contribution to drug metabolism,” said Michael Zimmermann, postdoctoral fellow in the Goodman lab and co-lead author of the study. “We think these approaches could shed light on how the gut microbiome also modulates our response to non-drug compounds, such as dietary nutrients and environmental agents.”
Funding: The work was primarily supported by the National Institutes of Health.
Source:
Yale
Media Contacts:
Bill Hathaway – Yale
Image Source:
The image is credited to Marieke F. Buffing.
Original Research: Closed access
“Mapping human microbiome drug metabolism by gut bacteria and their genes”. Michael Zimmermann, Maria Zimmermann-Kogadeeva, Rebekka Wegmann & Andrew L. Goodman.
Nature. doi:10.1038/s41586-019-1291-3
Abstract
Mapping human microbiome drug metabolism by gut bacteria and their genes
Individuals vary widely in drug responses, which can be dangerous and expensive due to treatment delays and adverse effects. Growing evidence implicates the gut microbiome in this variability, however the molecular mechanisms remain largely unknown. Here we measured the ability of 76 diverse human gut bacteria to metabolize 271 oral drugs and found that many of these drugs are chemically modified by microbes. We combined high-throughput genetics with mass spectrometry to systematically identify drug-metabolizing microbial gene products. These microbiome-encoded enzymes can directly and significantly impact intestinal and systemic drug metabolism in mice and can explain drug-metabolizing activities of human gut bacteria and communities based on their genomic contents. These causal links between microbiota gene content and metabolic activities connect interpersonal microbiome variability to interpersonal differences in drug metabolism, which has implications for medical therapy and drug development across multiple disease indications.