Summary: A new neuroimaging study finds many Alzheimer’s patients and those at risk of developing the disease show a combination of atrophy factors.
Source: Mass General.
Imaging studies imply that most patients, at-risk individuals show a combination of atrophy factors.
Mathematical modeling of the brain scans of patients with Alzheimer’s disease and others at risk for the devastating neurodegenerative disorder has identified specific patterns of brain atrophy that appear to be related to the loss of particular cognitive abilities. In their report that has been published online in the Proceedings of the National Academy of Sciences, a team of researchers at Massachusetts General Hospital (MGH) and the National University of Singapore describe how different atrophy patterns may explain the different ways that Alzheimer’s disease can be manifested in individual patients.
“The symptom severity and neurodegeneration can vary widely across patients in Alzheimer’s disease,” says Thomas Yeo, PhD, of the Martinos Center for Biomedical Imaging at MGH. “Our work shows that participants in this study exhibit at least three atrophy patterns – cortical, temporal or subcortical – that are associated with variability in cognitive decline not only in patients diagnosed with Alzheimer’s but also in individuals with mild cognitive impairment or those who are cognitively normal but are at risk for Alzheimer’s.” In addition to his affiliation with the Martinos Center, Yeo is an assistant professor in the Department of Electrical and Computer Engineering, Clinical Imaging Research Centre and Singapore Institute for Neurotechnology at the National University of Singapore.
The study analyzed data collected as part of the Alzheimer’s Disease Neuroimaging Initiative (ADNI), a multi-institutional project to develop biomarkers – including blood tests, cerebrospinal fluid tests, and imaging studies – that can be used for diagnosis or in clinical trials. Yeo and his team – including investigators at the MGH and in Singapore – analyzed MR images taken of the brains of 378 ADNI participants when they enrolled in the study. Of these participants, 188 had been diagnosed with Alzheimer’s disease; the others – 147 with mild cognitive impairment and 43 who were cognitively normal – were at increased risk based on levels in their brains of the beta-amyloid plaques that are characteristic of the disease.
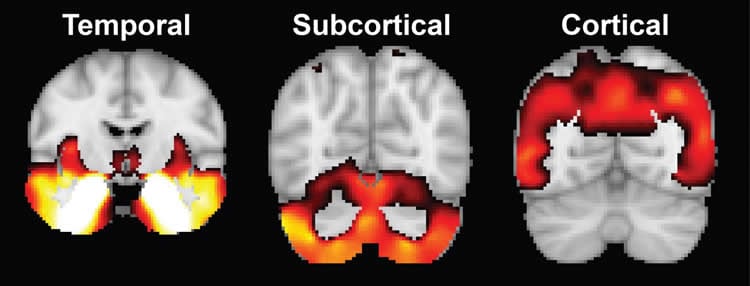
As a first step, the research team analyzed data from the baseline structural MRIs using a mathematical model that estimated the probability that particular details of each image were associated with atrophy of a specific location within the brain. Based on the location of atrophy factors, they determined three atrophy factor patterns: cortical – representing atrophy in most of the cerebral cortex; temporal – indicating atrophy in the temporal cortex (the cortical lobe behind the ears), hippocampus and amygdala; and subcortical, indicating atrophy in the cerebellum, striatum and thalamus, structures at the base of the brain.
Analysis of study participant scans taken two years later indicated that atrophy factor patterns were persistent in individuals and did not reflect different stages of disease. Most participants – including those in the mild cognitive impairment and cognitively normal groups – showed levels of more than one atrophy factor.
Behavioral and cognitive tests of study participants taken at six-month intervals indicated associations between particular atrophy factor patterns and specific cognitive deficits. Individuals in whom temporal atrophy predominated had greater problems with memory, while cortical atrophy was associated with difficulties with executive function – the ability to plan and to accomplish goals. Individual differences in how atrophy factors are distributed within the brain may allow prediction of the rate at which cognitive abilities would be expected to decline.
“Most previous studies focused on patients already diagnosed, but we were able to establish distinct atrophy patterns not only in diagnosed patients but also in at-risk participants who had mild impairment or were cognitively normal at the outset of the study,” Yeo says. “That is important because the neurodegenerative cascade that leads to Alzheimer’s starts years, possibly decades, before diagnosis. So understanding different atrophy patterns among at-risk individuals is quite valuable.
He adds, “Previous studies assumed that an individual can only express a single neurodegenerative pattern, which is highly restrictive since in any aged person there could be multiple pathological factors going on at the same time – such as vascular impairment along with the amyloid plaques and tau tangles that are directly associated with Alzheimer’s. So individuals who are affected by multiple, co-existing pathologies might be expected to exhibit multiple atrophy patterns.”
Future research could further determine whether and how these atrophy patterns relate to the distribution of amyloid and tau and the mechanisms by which they affect specific cognitive abilities, Yeo explains. The same analytic approach also could be applied to other types of patient data and extended to other neurodegenerative disorder that produce varying symptom patterns, such as Parkinson’s disease and autism.
Xiuming Zhang, of Yeo’s National University of Singapore lab, is lead author of the PNAS article. Additional co-authors are Elizabeth Mormino, PhD, and Reisa Sperling, MD, MGH Department of Neurology; Mert Sabuncu, PhD, Martinos Center, and Nanbo Sun, National University of Singapore.
Funding: The study was supported by grants from the National University of Singapore, Singapore National Medical Research Council and U.S. National Institutes of Health grants 1K25 EB013649-01, 1R21 AG050122-01A1, P01 AG036694, and F32AG044054.
Source: Terri Ogan – Mass General
Image Source: NeuroscienceNews.com image is credited to Xiuming Zhang, National University of Singapore.
Original Research: Full open access research for “Bayesian model reveals latent atrophy factors with dissociable cognitive trajectories in Alzheimer’s disease” by Xiuming Zhang, Elizabeth C. Mormino, Nanbo Sun, Reisa A. Sperling, Mert R. Sabuncu, B. T. Thomas Yeo, and the Alzheimer’s Disease Neuroimaging Initiative in PNAS. Published online October 4 2016 doi:10.1073/pnas.1611073113
[cbtabs][cbtab title=”MLA”]Mass General. “Different Brain Atrophy Patterns May Explain Variability in Alzheimer’s Symptoms.” NeuroscienceNews. NeuroscienceNews, 7 October 2016.
<https://neurosciencenews.com/alzheimers-atrophy-patterns-5237/>.[/cbtab][cbtab title=”APA”]Mass General. (2016, October 7). Different Brain Atrophy Patterns May Explain Variability in Alzheimer’s Symptoms. NeuroscienceNews. Retrieved October 7, 2016 from https://neurosciencenews.com/alzheimers-atrophy-patterns-5237/[/cbtab][cbtab title=”Chicago”]Mass General. “Different Brain Atrophy Patterns May Explain Variability in Alzheimer’s Symptoms.” https://neurosciencenews.com/alzheimers-atrophy-patterns-5237/ (accessed October 7, 2016).[/cbtab][/cbtabs]
Abstract
Bayesian model reveals latent atrophy factors with dissociable cognitive trajectories in Alzheimer’s disease
We used a data-driven Bayesian model to automatically identify distinct latent factors of overlapping atrophy patterns from voxelwise structural MRIs of late-onset Alzheimer’s disease (AD) dementia patients. Our approach estimated the extent to which multiple distinct atrophy patterns were expressed within each participant rather than assuming that each participant expressed a single atrophy factor. The model revealed a temporal atrophy factor (medial temporal cortex, hippocampus, and amygdala), a subcortical atrophy factor (striatum, thalamus, and cerebellum), and a cortical atrophy factor (frontal, parietal, lateral temporal, and lateral occipital cortices). To explore the influence of each factor in early AD, atrophy factor compositions were inferred in beta-amyloid–positive (Aβ+) mild cognitively impaired (MCI) and cognitively normal (CN) participants. All three factors were associated with memory decline across the entire clinical spectrum, whereas the cortical factor was associated with executive function decline in Aβ+ MCI participants and AD dementia patients. Direct comparison between factors revealed that the temporal factor showed the strongest association with memory, whereas the cortical factor showed the strongest association with executive function. The subcortical factor was associated with the slowest decline for both memory and executive function compared with temporal and cortical factors. These results suggest that distinct patterns of atrophy influence decline across different cognitive domains. Quantification of this heterogeneity may enable the computation of individual-level predictions relevant for disease monitoring and customized therapies. Factor compositions of participants and code used in this article are publicly available for future research.
“Bayesian model reveals latent atrophy factors with dissociable cognitive trajectories in Alzheimer’s disease” by Xiuming Zhang, Elizabeth C. Mormino, Nanbo Sun, Reisa A. Sperling, Mert R. Sabuncu, B. T. Thomas Yeo, and the Alzheimer’s Disease Neuroimaging Initiative in PNAS. Published online October 4 2016 doi:10.1073/pnas.1611073113






