A study of changes in the patterns of gene activity in the brains of people with Down syndrome reveals that the formation of the brain’s white matter is affected throughout life, a finding that suggests treatment might be possible for the condition that affects 400,000 Americans.
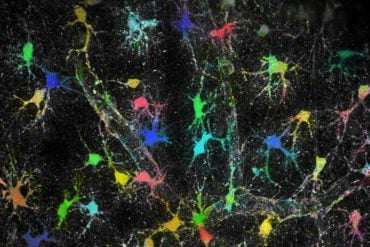
Researchers at Yale University and Boston University found abnormal patterns of gene expression in the developing brain of individuals with Down syndrome that influence the maturation of oligodendrocytes — the cells that make the fatty myelin coating that insulates nerve fibers and make rapid transmission of nerve impulses possible. These expression patterns continue into adulthood and are not limited to early development as previously believed, according to the study published Feb. 25 in the journal Neuron.
“Defective myelination may be one of the factors causing the intellectual disability that is a hallmark of Down syndrome,” said John Silbereis, postdoctoral fellow in the Department of Neuroscience and the Kavli Institute for Neuroscience at Yale, and co-first author of the paper.

It may be possible that individuals with Down syndrome might benefit from myelin-regenerating drugs under development to treat conditions like multiple sclerosis, which are also marked by altered myelination of nerve cells, the authors say.
Typically, people are all born with pairs of 23 chromosomes but all people with Down syndrome have a third copy of chromosome 21. However, the exact mechanisms that cause intellectual disabilities have been difficult to pin down. Under the direction of senior authors Nenad Sestan, professor of neuroscience, genetics and psychiatry at Yale, and Tarik F. Haydar at Boston University, researchers did a comprehensive genomic analysis of brain tissue across ages — from prenatal to 40-year-old adults. This analysis identified over 1,300 genes that were disrupted at some point in the development of individuals with Down syndrome. They found 95% of these genes were in regions of the genome other than chromosome 21.
These results indicate that triplication of chromosome 21 have large and widespread effects on gene expression patterns in individuals with Down syndrome, say the scientists.
Hyo Jung Kang of Yale is a co-first author of the paper. Sestan also holds posts at the Kavli Institute, Section of Comparative Medicine, Program in Cellular Neuroscience, Neurodegeneration and Repair, and the Child Study Center.
Source: Bill Hathaway – Yale
Image Source: The image is adapted from the Yale press release.
Original Research: Abstract for “Down Syndrome Developmental Brain Transcriptome Reveals Defective Oligodendrocyte Differentiation and Myelination” by Jose Luis Olmos-Serrano, Hyo Jung Kang, William A. Tyler, John C. Silbereis, Feng Cheng, Ying Zhu, Mihovil Pletikos, Lucija Jankovic-Rapan, Nathan P. Cramer, Zygmunt Galdzicki, Joseph Goodliffe, Alan Peters, Claire Sethares, Ivana Delalle, Jeffrey A. Golden, Tarik F. Haydar, and Nenad Sestan in Neuron. Published online February 25 2016 doi:10.1016/j.neuron.2016.01.042
Abstract
Down Syndrome Developmental Brain Transcriptome Reveals Defective Oligodendrocyte Differentiation and Myelination
Highlights
•Genome-wide spatiotemporal dysregulation of gene expression is found in DS brains
•Transcriptome changes reflect altered oligodendrocyte development and myelination
•Oligodendrocyte differentiation and myelination is altered in DS model mice (Ts65Dn)
•Speed of action potential propagation is decreased in Ts65Dn neocortical white matter
Summary
Trisomy 21, or Down syndrome (DS), is the most common genetic cause of developmental delay and intellectual disability. To gain insight into the underlying molecular and cellular pathogenesis, we conducted a multi-region transcriptome analysis of DS and euploid control brains spanning from mid-fetal development to adulthood. We found genome-wide alterations in the expression of a large number of genes, many of which exhibited temporal and spatial specificity and were associated with distinct biological processes. In particular, we uncovered co-dysregulation of genes associated with oligodendrocyte differentiation and myelination that were validated via cross-species comparison to Ts65Dn trisomy mice. Furthermore, we show that hypomyelination present in Ts65Dn mice is in part due to cell-autonomous effects of trisomy on oligodendrocyte differentiation and results in slower neocortical action potential transmission. Together, these results identify defects in white matter development and function in DS, and they provide a transcriptional framework for further investigating DS neuropathogenesis.
“Down Syndrome Developmental Brain Transcriptome Reveals Defective Oligodendrocyte Differentiation and Myelination” by Jose Luis Olmos-Serrano, Hyo Jung Kang, William A. Tyler, John C. Silbereis, Feng Cheng, Ying Zhu, Mihovil Pletikos, Lucija Jankovic-Rapan, Nathan P. Cramer, Zygmunt Galdzicki, Joseph Goodliffe, Alan Peters, Claire Sethares, Ivana Delalle, Jeffrey A. Golden, Tarik F. Haydar, and Nenad Sestan in Neuron. Published online February 25 2016 doi:10.1016/j.neuron.2016.01.042