Summary: Study goes beyond evaluating the organisms in the microbiome, looking at the functions different bacteria may be performing.
Source: Oregon State University
Researchers at Oregon State University have made an important advance in understanding the roles that gut bacteria play in human health.
Learning the mechanisms by which gut microbes affect the health of their hosts opens the door to the development of better, more personalized diagnostic methods and therapies.
Most studies so far have focused on how the composition of the microbiome — i.e., which organisms are present, and in what amounts — associates with health in general or various diseases.
The OSU research led by Ph.D. student Courtney Armour goes a step further by looking not just at which organisms are in the microbiome, but also what functions they might be performing. Findings were published in mSystems.
Armour, working under microbiology and statistics researcher Thomas Sharpton in OSU’s College of Science, analyzed data and findings from eight different studies encompassing seven different diseases in a metagenomic meta-analysis.
Metagenomics refers to the study of genetic material recovered directly from environmental samples — in this case, human fecal samples — as opposed to from organisms cultured in a lab. A meta-analysis is a statistical technique for combining data from multiple studies.
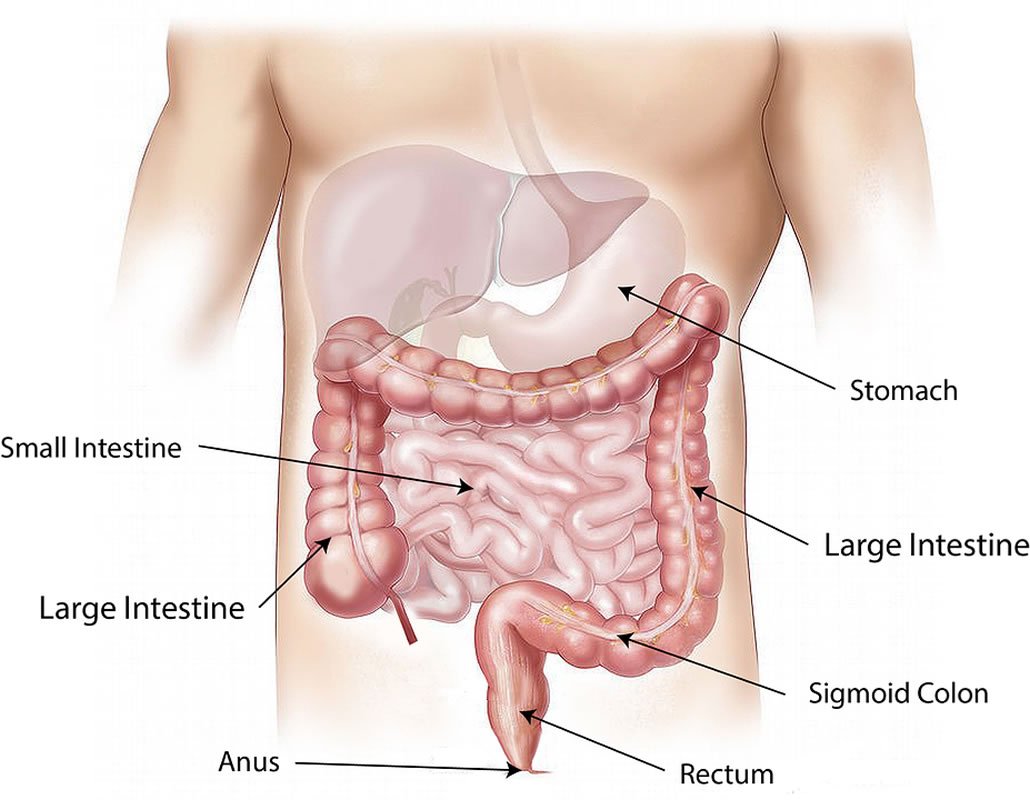
The meta-analysis performed by Armour, Sharpton and their collaborators involved metagenomic data from nearly 2,000 stool samples collected for studies involving colorectal cancer, Crohn’s disease, liver cirrhosis, obesity, rheumatoid arthritis, type 2 diabetes and ulcerative colitis.
The gut microbiota features more than 10 trillion microbial cells from about 1,000 different bacterial species. The microbial ecosystem stays in balance via cell-to-cell signaling and the release of antimicrobial peptides that keep in check certain bacterial clades.
Gut microbes interact with their human host as well, sometimes in ways that promote health, other times in ways that contribute to disease development. Dysbiosis, or imbalance, in the microbiome is commonly associated with detrimental effects to the host’s health.
“In our study, we looked at how gut microbiome protein family richness, composition and dispersion relate to disease,” Sharpton said.
Proteins are large, complex molecules that do most of the work in cells and are required for the structure, function and regulation of tissues and organs.
“Our analysis of protein family richness showed that patients with Crohn’s disease, obesity, type 2 diabetes or ulcerative colitis feature a smaller number of protein families compared to their respective control populations,” Sharpton said. “On the other hand, people with colorectal cancer had a larger number of microbiome protein families than their controls.”
The researchers also looked at “beta-dispersion,” which measures the compositional variation of the microbiome among a group of individuals.
“Prior work linked disease to an increase in taxonomic beta-dispersion,” Sharpton said. “We looked at whether gut microbiome functional beta-dispersion is different between healthy and diseased populations and saw an increase in functional beta-dispersion in patients with colorectal cancer, Crohn’s disease and liver cirrhosis. Individuals with obesity displayed reduced beta-dispersion relative to their controls.”
The amount of overlap — functions associating with multiple diseases — was striking, said Armour, who added there’s much more to learn.
“We really need more data,” she said. “And we need more information about the subjects in the studies, about other things that may be affecting the microbiome, things like diet and geography. We need more data from diverse locations and populations to account for sources of variations.”
The long-term goal, Sharpton said, is for doctors to be able to use information derived from metagenomics to diagnose diseases “more specifically, more quickly and less invasively.”
“Our work points to information coded in the metagenome that could be used for that, but that requires more data to make those diagnoses more robust,” he said. “We’re trying to disentangle cause and effect, to resolve these needles in haystacks and find the links between the microbiome and health. Future research can leverage this new knowledge to test microbiome functions against the presence and severity of various diseases.”
Funding: The National Institutes of Health and the National Science Foundation supported this research, which included scientists from the Lawrence Berkeley National Laboratory, the University of California, San Francisco, and Gladstone Institutes.
Source:
Oregon State University
Media Contacts:
Steve Lundeberg – Oregon State University
Image Source:
The image is in the public domain.
Original Research: Open access
“A Metagenomic Meta-analysis Reveals Functional Signatures of Health and Disease in the Human Gut Microbiome”. Courtney R. Armour, Stephen Nayfach, Katherine S. Pollard, Thomas J. Sharpton.
mSystems. doi:10.1128/mSystems.00332-18
Abstract
A Metagenomic Meta-analysis Reveals Functional Signatures of Health and Disease in the Human Gut Microbiome
While recent research indicates that human health is affected by the gut microbiome, the functional mechanisms that underlie host-microbiome interactions remain poorly resolved. Metagenomic clinical studies can address this problem by revealing specific microbial functions that stratify healthy and diseased individuals. To improve our understanding of the relationship between the gut microbiome and health, we conducted the first integrative functional analysis of nearly 2,000 publicly available fecal metagenomic samples obtained from eight clinical studies. We identified characteristics of the gut microbiome that associate generally with disease, including functional alpha-diversity, beta-diversity, and beta-dispersion. Using regression modeling, we identified specific microbial functions that robustly stratify diseased individuals from healthy controls. Many of these functions overlapped multiple diseases, suggesting a general role in host health, while others were specific to a single disease and may indicate disease-specific etiologies. Our results clarify potential microbiome-mediated mechanisms of disease and reveal features of the microbiome that may be useful for the development of microbiome-based diagnostics.
IMPORTANCE The composition of the gut microbiome associated with a wide range of human diseases, but the mechanisms underpinning these associations are not well understood. To shift toward a mechanistic understanding, we integrated distinct metagenomic data sets to identify functions encoded in the gut microbiome that associate with multiple diseases, which may be important to human health. Additionally, we identified functions that associate with specific diseases, which may elucidate disease-specific etiologies. We demonstrated that the functions encoded in the microbiome can be used to classify disease status, but the inclusion of additional patient covariates may be necessary to obtain sufficient accuracy. Ultimately, this analysis advances our understanding of the gut microbiome functions that constitute a healthy microbiome and identifies potential targets for microbiome-based diagnostics and therapeutics.