Summary: Dysregulation of Zfp36l1 causes neural lineage progenitor cells to grow rapidly, taking on the properties of fast growing stem cells as they become cancerous.
Source: Cincinnati Children’s Hospital Medical Center
Cell-by-cell genetic analyses of developing brain tissues in neonatal mice and laboratory models of brain cancer allowed scientists to discover a molecular driver of the highly aggressive, deadly, and treatment-resistant brain cancer, glioblastoma.
Published findings in Cell Stem Cell describe how the single-cell analyses identified a subpopulation of cells critical to glioblastoma formation–the early primitive progenitor cells of oligodendrocyte brain cells, pri-OPC progenitors, according to Q. Richard Lu, PhD, lead investigator and Scientific Director of the Brain Tumor Center at Cincinnati Children’s Hospital Medical Center.
The data suggest that reprogramming of primitive oligodendrocyte progenitors into a stem-like state plays an important role in glioma initiation and progression. The researchers’ primary molecular target in the study, a protein called Zfp36l1, launches biological programs that mirror those of healthy early brain development in the mice, but instead help fuel brain cancer growth. The discovery presents an opportunity to find out if new therapeutic approaches can stop glioblastoma at its earliest stages of initial formation or recurrence, Lu said.
Researchers learned that dysregulation of Zfp36l1–a protein that binds together different RNAs–causes neural lineage progenitor cells to grow so rapidly that they take on the properties of fast-growing stem-like cells as they turn cancerous. This was especially true of less-developed primitive oligodendrocyte progenitor cells. The discovery gave researchers an idea.
“When we used small RNA-interfering molecules to inhibit the expression of the Zpf36l1 gene in mouse and human glioma tumor cells, it significantly decreased the rate of glioma cell growth and spread,” Lu said. “In mice with glioblastoma tumors, genetic knockout of Zpf36l1 significantly increased survival time for the animals.”
Single-Cell Transcriptomics
The researchers used single-cell transcriptomics as a foundation for their study. This allowed them to identify, and compare, all of the RNA molecules in each developing glial brain cell and developing cancerous glioblastoma cell. This included the distinct populations of transitioning oligodendrocyte and astrocyte progenitor cells, particularly the early pri-OPC progenitors that support the rapid growth of glioblastoma.
Researchers said this study is important because prior to the development of new technologies that allow for single-cell analysis, the extremely diverse genetics and types of brain cells that cause high-grade glioblastoma have been hard to nail down.
To determine if their observations in mouse glioma models might apply to tumor formation in human brain tumors, researchers also analyzed single-cell transcriptomics of human tumor cells donated by patients. They found similar populations and genetic signatures in human oligodendrocyte progenitor cells. The cells had regressed to a less-developed mitotic state and assumed the characteristics of fast-growing stem-like brain cells, setting the stage for tumorigenesis.
Preclinical Research Ongoing
The researchers emphasized their data were produced in mice and other laboratory models of brain cancer, so it still remains unknown if the results will translate to patient treatment. The study does provide important clues, however, for battling a cancer that usually is fatal, including in young children, researchers said.
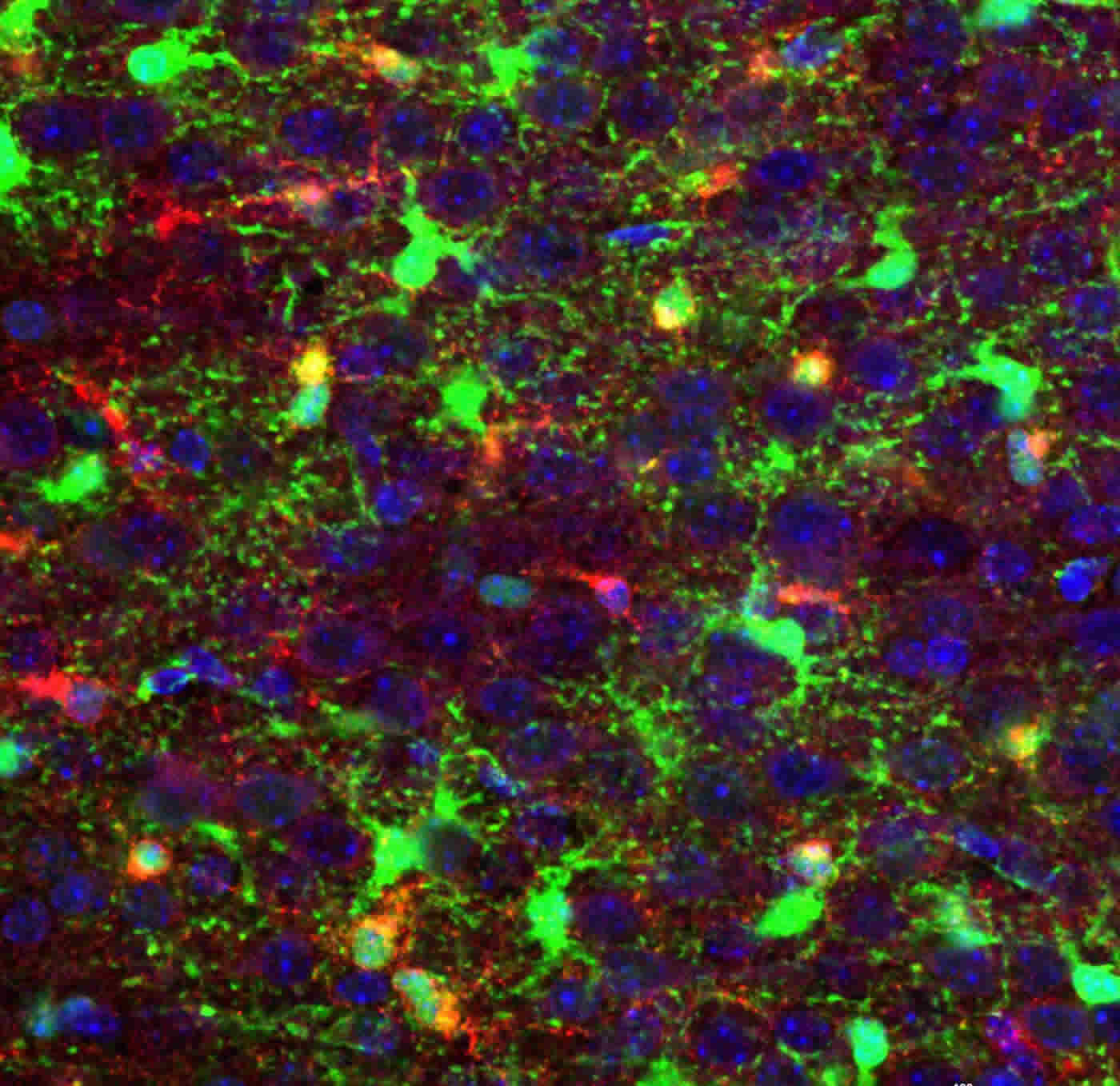
Now that Lu and his colleagues know blocking Zfp36l1 function in glioma cells forming from primitive oligodendrocyte progenitor cells inhibits glioblastoma growth in lab models, they are expanding their efforts. They want to find an effective way to therapeutically exploit what they consider to be a potential new vulnerability for a disease that so far has been invincible. This includes future work to develop specific therapeutic agents that might more precisely and effectively target Zfp36l1 and the brain cells it helps turn cancerous.
The study’s first author is Qinjie Weng, PhD, a former member of the Lu laboratory and now an associate professor at Zhejiang University in China. The research team was comprised of researchers at Cincinnati Children’s and collaborators from seven other research institutions in the United States and China.
Funding: Funding support came in part from the U.S. National Institutes of Health (R37NSO75243, RO1NSO75243) and the Intramural Research Program of the National Institute of Environmental Health Sciences.
Source:
Cincinnati Children’s Hospital Medical Center
Media Contacts:
Nick Miller – Cincinnati Children’s Hospital Medical Center
Image Source:
The image is credited to Cincinnati Children’s.
Original Research: Closed access.
Q. Richard Lu, et al. “Single-Cell Transcriptomics Uncovers Glial Progenitor Diversity and Cell Fate Determinants during Development and Gliomagenesis”. Cell Stem Cell 2019;68(No. SS-2):1–19. doi:10.1016/j.stem.2019.03.006
Abstract
Single-Cell Transcriptomics Uncovers Glial Progenitor Diversity and Cell Fate Determinants during Development and Gliomagenesis
Highlights
• scRNA-seq reveals distinct transitional states of glial progenitors in the cortex
• Intersectional analysis reveals malignant analogs of glial progenitor intermediates
• Reprogramming OPC intermediates into a stem-like state for malignant gliomagenesis
• Zfp36l1 is a critical regulator of glial fate specification and gliomagenesis
Summary
The identity and degree of heterogeneity of glial progenitors and their contributions to brain tumor malignancy remain elusive. By applying lineage-targeted single-cell transcriptomics, we uncover an unanticipated diversity of glial progenitor pools with unique molecular identities in developing brain. Our analysis identifies distinct transitional intermediate states and their divergent developmental trajectories in astroglial and oligodendroglial lineages. Moreover, intersectional analysis uncovers analogous intermediate progenitors during brain tumorigenesis, wherein oligodendrocyte-progenitor intermediates are abundant, hyper-proliferative, and progressively reprogrammed toward a stem-like state susceptible to further malignant transformation. Similar actively cycling intermediate progenitors are prominent components in human gliomas with distinct driver mutations. We further unveil lineage-driving networks underlying glial fate specification and identify Zfp36l1 as necessary for oligodendrocyte-astrocyte lineage transition and glioma growth. Together, our results resolve the dynamic repertoire of common and divergent glial progenitors during development and tumorigenesis and highlight Zfp36l1 as a molecular nexus for balancing glial cell-fate decision and controlling gliomagenesis.