Summary: Researchers use gene editing technology to help partially restore vision to blind rodents.
Source: Salk Institute.
Salk researchers have discovered, for the first time, how to place DNA in specific locations in non-dividing cells.
Salk Institute researchers have discovered a holy grail of gene editing—the ability to, for the first time, insert DNA at a target location into the non-dividing cells that make up the majority of adult organs and tissues. The technique, which the team showed was able to partially restore visual responses in blind rodents, will open new avenues for basic research and a variety of treatments, such as for retinal, heart and neurological diseases.
“We are very excited by the technology we discovered because it’s something that could not be done before,” says Juan Carlos Izpisua Belmonte, a professor in Salk’s Gene Expression Laboratory and senior author of the paper published on November 16, 2016 in Nature. “For the first time, we can enter into cells that do not divide and modify the DNA at will. The possible applications of this discovery are vast.”
Until now, techniques that modify DNA—such as the CRISPR-Cas9 system—have been most effective in dividing cells, such as those in skin or the gut, using the cells’ normal copying mechanisms. The new Salk technology is ten times more efficient than other methods at incorporating new DNA into cultures of dividing cells, making it a promising tool for both research and medicine. But, more importantly, the Salk technique represents the first time scientists have managed to insert a new gene into a precise DNA location in adult cells that no longer divide, such as those of the eye, brain, pancreas or heart, offering new possibilities for therapeutic applications in these cells.
To achieve this, the Salk researchers targeted a DNA-repair cellular pathway called NHEJ (for “non-homologous end-joining”), which repairs routine DNA breaks by rejoining the original strand ends. They paired this process with existing gene-editing technology to successfully place new DNA into a precise location in non-dividing cells.
“Using this NHEJ pathway to insert entirely new DNA is revolutionary for editing the genome in live adult organisms,” says Keiichiro Suzuki, a senior research associate in the Izpisua Belmonte lab and one of the paper’s lead authors. “No one has done this before.”
First, the Salk team worked on optimizing the NHEJ machinery for use with the CRISPR-Cas9 system, which allows DNA to be inserted at very precise locations within the genome. The team created a custom insertion package made up of a nucleic acid cocktail, which they call HITI, or homology-independent targeted integration. Then they used an inert virus to deliver HITI’s package of genetic instructions to neurons derived from human embryonic stem cells.
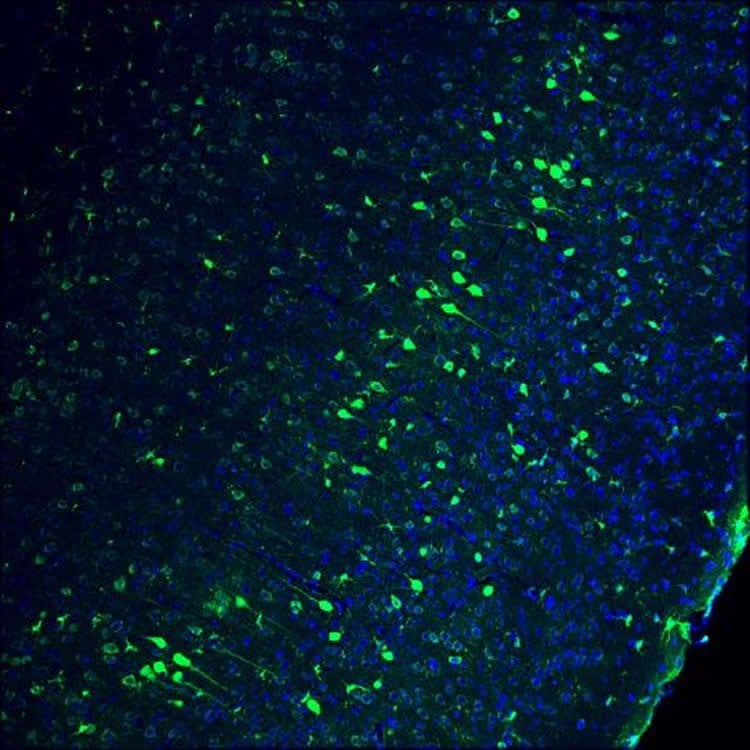
“That was the first indication that HITI might work in non-dividing cells,” says Jun Wu, staff scientist and co-lead author. With that feat under their belts, the team then successfully delivered the construct to the brains of adult mice. Finally, to explore the possibility of using HITI for gene-replacement therapy, the team tested the technique on a rat model for retinitis pigmentosa, an inherited retinal degeneration condition that causes blindness in humans. This time, the team used HITI to deliver to the eyes of 3-week-old rats a functional copy of Mertk, one of the genes that is damaged in retinitis pigmentosa. Analysis performed when the rats were 8 weeks old showed that the animals were able to respond to light, and passed several tests indicating healing in their retinal cells.
“We were able to improve the vision of these blind rats,” says co-lead author Reyna Hernandez-Benitez, a Salk research associate. “This early success suggests that this technology is very promising.”
The team’s next steps will be to improve the delivery efficiency of the HITI construct. As with all genome editing technologies, getting enough cells to incorporate the new DNA is a challenge. The beauty of HITI technology is that it is adaptable to any targeted genome engineering system, not just CRISPR-Cas9. Thus, as the safety and efficiency of these systems improve, so too will the usefulness of HITI.
“We now have a technology that allows us to modify the DNA of non-dividing cells, to fix broken genes in the brain, heart and liver,” says Izpisua Belmonte. “It allows us for the first time to be able to dream of curing diseases that we couldn’t before, which is exciting.”
Other researchers on the study were Euiseok J. Kim, Fumiyuki Hatanaka, Mako Yamamoto, Toshikazu Araoka, Masakazu Kurita, Tomoaki Hishida, Mo Li, Emi Aizawa, April Goebl, Rupa Devi Soligalla, Concepcion Rodriguez Esteban, Travis Berggren and Edward M. Callaway of the Salk Institute; Yuji Tsunekawa and Fumio Matsuzaki of RIKEN Center for Developmental Biology; Pierre Magistretti of King Abdullah University of Science and Technology; Jie Zhu, Tingshuai Jiang, Xin Fu, Maryam Jafari and Kang Zhang of Shiley Eye Institute and Institute for Genomic Medicine, University of California San Diego; Zhe Li, Shicheng Guo, Song Chen and Kun Zhang of Institute of Engineering in Medicine, University of California San Diego; Jing Qu and Guang-Hui Liu of Chinese Academy of Sciences; Jeronimo Lajara, Estrella Nuñez and Pedro Guillen of Universidad Catolica San Antonio de Murcia; and Josep M. Campistol of the University of Barcelona.
Funding: The work and the researchers involved were supported in part by the National Institutes of Health, The Leona M. and Harry B. Helmsley Charitable Trust, the G. Harold and Leila Y. Mathers Charitable Foundation, The McKnight Foundation, The Moxie Foundation, the Dr. Pedro Guillen Foundation and Universidad Catolica San Antonio de Murcia, Spain.
Source: Salk Institute
Image Source: NeuroscienceNews.com image is credited to Salk Institute.
Video Source: The video is credited to Salk Institute,
Original Research: Abstract for “In vivo genome editing via CRISPR/Cas9 mediated homology-independent targeted integration” by Keiichiro Suzuki, Yuji Tsunekawa, Reyna Hernandez-Benitez, Jun Wu, Jie Zhu, Euiseok J. Kim, Fumiyuki Hatanaka, Mako Yamamoto, Toshikazu Araoka, Zhe Li, Masakazu Kurita, Tomoaki Hishida, Mo Li, Emi Aizawa, Shicheng Guo, Song Chen, April Goebl, Rupa Devi Soligalla, Jing Qu, Tingshuai Jiang, Xin Fu, Maryam Jafari, Concepcion Rodriguez Esteban, W. Travis Berggren, Jeronimo Lajara, Estrella Nuñez-Delicado, Pedro Guillen, Josep M. Campistol, Fumio Matsuzaki, Guang-Hui Liu, Pierre Magistretti, Kun Zhang, Edward M. Callaway, Kang Zhang & Juan Carlos Izpisua Belmonte Nature. Published online November 16 2016 doi:10.1038/nature20565
[cbtabs][cbtab title=”MLA”]Salk Institute “New Gene-Editing Technology Partially Restores Vision in Blind Animals.” NeuroscienceNews. NeuroscienceNews, 16 November 2016.
<https://neurosciencenews.com/crispr-vision-restoration-5540/>.[/cbtab][cbtab title=”APA”]Salk Institute (2016, November 16). New Gene-Editing Technology Partially Restores Vision in Blind Animals. NeuroscienceNew. Retrieved November 16, 2016 from https://neurosciencenews.com/crispr-vision-restoration-5540/[/cbtab][cbtab title=”Chicago”]Salk Institute “New Gene-Editing Technology Partially Restores Vision in Blind Animals.” https://neurosciencenews.com/crispr-vision-restoration-5540/ (accessed November 16, 2016).[/cbtab][/cbtabs]
Abstract
In vivo genome editing via CRISPR/Cas9 mediated homology-independent targeted integration
Targeted genome editing via engineered nucleases is an exciting area of biomedical research and holds potential for clinical applications. Despite rapid advances in the field, in vivo targeted transgene integration is still infeasible because current tools are inefficient, especially for non-dividing cells, which compose most adult tissues. This poses a barrier for uncovering fundamental biological principles and developing treatments for a broad range of genetic disorders. Based on clustered regularly interspaced short palindromic repeat/Cas9 (CRISPR/Cas9) technology, here we devise a homology-independent targeted integration (HITI) strategy, which allows for robust DNA knock-in in both dividing and non-dividing cells in vitro and, more importantly, in vivo (for example, in neurons of postnatal mammals). As a proof of concept of its therapeutic potential, we demonstrate the efficacy of HITI in improving visual function using a rat model of the retinal degeneration condition retinitis pigmentosa. The HITI method presented here establishes new avenues for basic research and targeted gene therapies.
“In vivo genome editing via CRISPR/Cas9 mediated homology-independent targeted integration” by Keiichiro Suzuki, Yuji Tsunekawa, Reyna Hernandez-Benitez, Jun Wu, Jie Zhu, Euiseok J. Kim, Fumiyuki Hatanaka, Mako Yamamoto, Toshikazu Araoka, Zhe Li, Masakazu Kurita, Tomoaki Hishida, Mo Li, Emi Aizawa, Shicheng Guo, Song Chen, April Goebl, Rupa Devi Soligalla, Jing Qu, Tingshuai Jiang, Xin Fu, Maryam Jafari, Concepcion Rodriguez Esteban, W. Travis Berggren, Jeronimo Lajara, Estrella Nuñez-Delicado, Pedro Guillen, Josep M. Campistol, Fumio Matsuzaki, Guang-Hui Liu, Pierre Magistretti, Kun Zhang, Edward M. Callaway, Kang Zhang & Juan Carlos Izpisua Belmonte Nature. Published online November 16 2016 doi:10.1038/nature20565