NIH-funded study provides new platform for testing treatments for several neurodegenerative disorders.
Scientists at Mayo Clinic, Jacksonville, Florida created a novel mouse that exhibits the symptoms and neurodegeneration associated with the most common genetic forms of frontotemporal dementia (FTD) and amyotrophic lateral sclerosis (ALS, Lou Gehrig’s disease), both of which are caused by a mutation in the a gene called C9ORF72. The study was partially funded by the National Institutes of Health and published in the journal Science.
More than 30,000 Americans live with ALS, which destroys nerves that control essential movements, including speaking, walking, breathing and swallowing. After Alzheimer’s disease, FTD is the most common form of early onset dementia. It is characterized by changes in personality, behavior and language due to loss of neurons in the brain’s frontal and temporal lobes. Patients with mutations in the chromosome 9 open reading frame 72 (C9ORF72) gene have all or some symptoms associated with both disorders.
“Our mouse model exhibits the pathologies and symptoms of ALS and FTD seen in patients with the C9ORF72 mutation,” said the study’s lead author, Leonard Petrucelli, Ph.D., chair and Ralph and Ruth Abrams Professor of the Department of Neuroscience at Mayo Clinic, and a senior author of the study. “These mice could greatly improve our understanding of ALS and FTD and hasten the development of effective treatments.”
To create the model, Ms. Jeannie Chew, a Mayo Graduate School student and member of Dr. Petrucelli’s team, injected the brains of newborn mice with a disease-causing version of the C9ORF72 gene. As the mice aged, they became hyperactive, anxious, and antisocial, in addition to having problems with movement that mirrored patient symptoms. The brains of the mice were smaller than normal and had fewer neurons in areas that controlled the affected behaviors. The scientists also found that the mouse brains had key hallmarks of the disorders, including toxic clusters of ribonucleic acids (RNA) and TDP-43, a protein that has long been known to go awry in the majority of ALS and FTD cases.
“This is a significant advancement for the field. Scientists have been trying to create mice that accurately mimic the pathologies associated with these forms of ALS and FTD,” said Margaret Sutherland, Ph.D., program director, the National Institute of Neurological Disorders and Stroke, part of NIH. “This mouse model will be a valuable tool for developing therapies for these devastating disorders.”
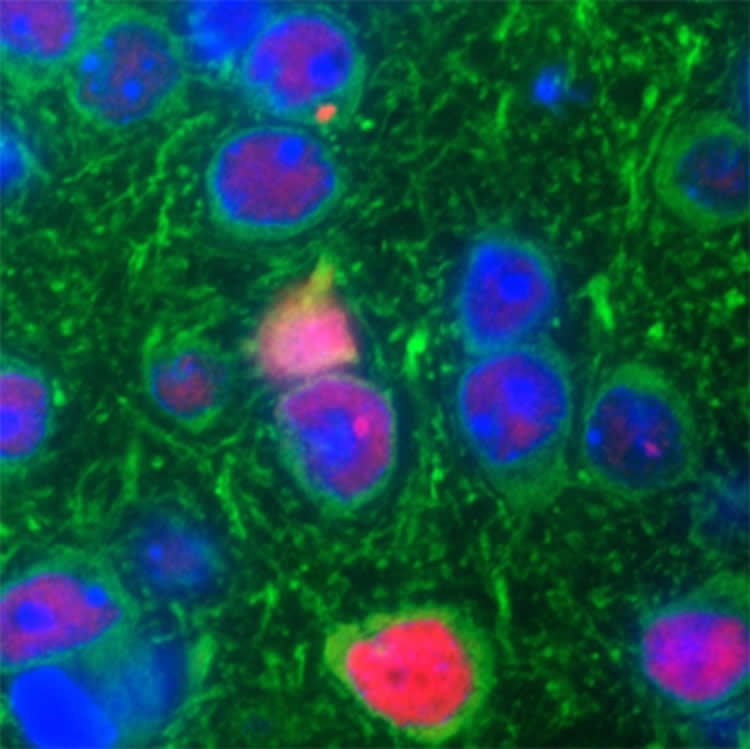
The C9ORF72 gene is encoded by repeating strings of six DNA molecules. Disease-causing C9ORF72 mutations make the strings excessively long which leads to the accumulation of RNA that either cluster into structures, called foci, or cause the production of abnormal c9RAN proteins in the brain and spinal cord of patients. The scientists found both in the brains of the mice. They also found clumps, or inclusions, of TDP-43 protein which is another pathological hallmark found in patients with the C9ORF72 mutation.
“Finding TDP-43 in these mice was unexpected” Dr. Petrucelli said. “We don’t yet know how foci and c9RAN proteins are linked to TDP-43 abnormalities, but with our new animal model, we now have a way to find out.”
Dr. Petrucelli and his team think these results are an important step in the development of therapies for these forms of ALS and FTD and other neurodegenerative disorders.
Funding: This work was supported by grants from NIH (NS089979, NS084528, NS079807, NS088689, NS063964, NS077402, NS084974, AG016574, ES20395), Department of Defense (ALSRP AL130125), Mayo Clinic Foundation, Mayo Clinic Center for Regenerative Medicine, Mayo Graduate School, ALS Association, Robert Packard Center for ALS Research at Johns Hopkins, Target ALS and Alzheimer’s Association (NIRP-14-304425, NIRP-12-259289).
Source: Christopher G. Thomas – NINDS/NIH
Image Source: Image is credited to Courtesy of Petrucelli lab, Mayo Clinic, Jacksonville, Fla.
Original Research: Abstract for “C9ORF72 repeat expansions in mice cause TDP-43 pathology, neuronal loss, and behavioral deficits” by Jeannie Chew, Tania F. Gendron, Mercedes Prudencio, Hiroki Sasaguri, Yong-Jie Zhang, Monica Castanedes-Casey, Chris W. Lee, Karen Jansen-West, Aishe Kurti, Melissa E. Murray, Kevin F. Bieniek, Peter O. Bauer, Ena C. Whitelaw, Linda Rousseau, Jeannette N. Stankowski, Caroline Stetler, Lillian M. Daughrity, Emilie A. Perkerson, Pamela Desaro, Amelia Johnston, Karen Overstreet, Dieter Edbauer, Rosa Rademakers, Kevin B. Boylan, Dennis W. Dickson, John D. Fryer, and Leonard Petrucelli in Science. Published online May 14 2015 doi:10.1126/science.aaa9344
Abstract
C9ORF72 repeat expansions in mice cause TDP-43 pathology, neuronal loss, and behavioral deficits
The major genetic cause of frontotemporal dementia and amyotrophic lateral sclerosis is a G4C2 repeat expansion in C9ORF72. Efforts to combat neurodegeneration associated with “c9FTD/ALS” are hindered by a lack of animal models recapitulating disease features. We developed a mouse model to mimic both neuropathological and clinical c9FTD/ALS phenotypes. We expressed (G4C2)66 throughout the murine central nervous system by means of somatic brain transgenesis mediated by adeno-associated virus. Brains of 6-month-old mice contained nuclear RNA foci, inclusions of poly(Gly-Pro), poly(Gly-Ala), and poly(Gly-Arg) dipeptide repeat proteins, as well as TDP-43 pathology. These mouse brains also exhibited cortical neuron and cerebellar Purkinje cell loss, astrogliosis, and decreased weight. (G4C2)66-mice also developed behavioral abnormalities similar to clinical symptoms of c9FTD/ALS patients, including hyperactivity, anxiety, antisocial behavior, and motor deficits.
“C9ORF72 repeat expansions in mice cause TDP-43 pathology, neuronal loss, and behavioral deficits” by Jeannie Chew, Tania F. Gendron, Mercedes Prudencio, Hiroki Sasaguri, Yong-Jie Zhang, Monica Castanedes-Casey, Chris W. Lee, Karen Jansen-West, Aishe Kurti, Melissa E. Murray, Kevin F. Bieniek, Peter O. Bauer, Ena C. Whitelaw, Linda Rousseau, Jeannette N. Stankowski, Caroline Stetler, Lillian M. Daughrity, Emilie A. Perkerson, Pamela Desaro, Amelia Johnston, Karen Overstreet, Dieter Edbauer, Rosa Rademakers, Kevin B. Boylan, Dennis W. Dickson, John D. Fryer, and Leonard Petrucelli in Science. Published online May 14 2015 doi:10.1126/science.aaa9344






