For Harvard neurobiologist Jeff Lichtman, the question hasn’t been whether scientists will ever understand the brain, but how closely they’ll have to look before they do.
The answer, it turns out, is very, very close.
Led by Lichtman, the Jeremy R. Knowles Professor of Molecular and Cellular Biology and Santiago Ramón y Cajal Professor of Arts and Sciences, a team of researchers has succeeded in comprehensively imaging — at the nano scale — a small portion of mouse brain. What they found, Lichtman said, could open the door to understanding how learning alters the brain.
A team of scientists from universities including Harvard, Johns Hopkins, and MIT contributed to the research by helping build the imaging acquisition and analysis pipeline required to study the brain in such detail. The co-authors are Narayanan Kasthuri, Kenneth Hayworth, Daniel Berger, Richard Schalek, José Angel Conchello, Seymour Knowles-Barley, Dongil Lee, Amelio Vázquez-Reina, Verena Kaynig, Thouis Jones, Mike Roberts, Josh Morgan, Juan Carlos Tapia, H. Sebastian Seung, William Gray Roncal, Joshua Vogelstein, Randal Burns, Daniel Sussman, Carey Priebe, and Hanspeter Pfister.
“One thing that surprised us was … that axons often made two, three, or even more synapses on the same dendrite,” Lichtman said. “The understanding had been that dendritic spines are there to collect information from as many different axons as possible, yet we found many cases where the same axon found different spines on the same dendrite.
“That was interesting because it suggests those multiple-contact axons are communicating more powerfully with the dendrite because they have more connections with it,” he continued. “When we looked closely, we found that it’s not possible for this result to happen by chance — some axons preferred to form synapses on some nearby dendrites more than they did with other equally nearby dendrites. That says that even in this trivially small volume, we are already beginning to see that the brain’s wiring diagram is organized in some interesting ways. It is not simply that nerve cells establish random contacts with nearby neurons, as was expected. There’s something purposeful going on here.”
The study also suggested that dendritic spines are not shaped by axons’ electrical activity, contrary to wide belief.
“The shape of spines goes from long and skinny to very short and stubby,” Lichtman said. “It’s thought that electrical activity, based on the axon, should generate that shape, but because we had multiple spines with different shapes enervated by the same axon, we know they have the same electrical activity.”
For Lichtman and his colleagues, the study is the culmination of years of effort, not only to understand the brain, but also to develop systems for collecting precise images of how the brain is wired.
“It’s a very, very tall staircase that we are trying to climb, but at least we’re on the staircase,” Lichtman said.
Getting to step one was a yearlong process.
“This paper took somewhere between five and six years to complete,” Lichtman said. “Much of that time was devoted to inventing the pipeline we use to capture these images. That required developing a means of cutting brain very thin, collecting the brain sections, and using this tape-based method that had not been used previously.”

With the system in place, Lichtman and colleagues set about using electron microscopes to capture images of tissue and shaping a method to trace cells through the various layers, allowing researchers to reconstruct axons, dendrites, and synapses into 3-D images. The images were also used to build a database that was mined for insights into neuron connectivity.
“What this paper describes is the pipeline where we start with a physical piece of brain and end up at the other end with a digital brain,” Lichtman said. “It’s been digitized and is minable, so you can look at this digital brain sample over and over again without having to dissect a real brain for each new question.”
The important insights in the study required a surprisingly small amount of brain.
“We imaged a section of brain that was 40-by-40-by-40 microns, and in that volume we completely saturated and reconstructed an area that was only 1,500 cubic microns,” Lichtman said. “It’s about three-billionths the size of a mouse brain, so it’s very small, but you have to start somewhere.”
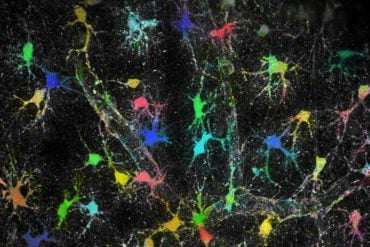
The selected region centers on a few dendritic processes of two large brain cells, allowing researchers to get a sense of just how many other nerve cells are located in the immediate vicinity of one small segment of a few neurons.
What they discovered came as a surprise even to Lichtman.
“We found 1,500 nerve cells provide nerve cell axons and dendrites in this little volume, which is a shockingly large number,” he said. “Those nerve cells contribute to a lot of other areas as well, but that gives you some sense of the extraordinary networking in the brains of mammals.”
Harvard University’s Jeff Lichtman and colleagues use automated EM technologies to probe the structures of neural tissue at nanometer resolution.
In fact, connections in the brain are so densely packed, state-of-the-art imaging can only begin to scratch the surface.
“We found a synapse approximately every cubic micron,” Lichtman said. “That means if you look at images of the brain captured using high-resolution techniques like functional magnetic resonance imaging, where the pixel size represents one cubic millimeter, then there are one billion synapses within every pixel.”
It may be there, in that staggering density, he said, that the study’s final message lies.
“It’s very hard to get a complete feeling for this,” Lichtman said. “There is this naïve view that if you just know a little bit more, then understanding it will be easier, but in this case, knowing a little more has shown us how much further we have to go before we’ll understand brains.”
Source: Peter Reuell – Harvard
Image Credit: The image is credited to Cell
Video Source: The video is available at the Cell Video Abstracts YouTube page
Original Research: Abstract for “Saturated Reconstruction of a Volume of Neocortex” by Jeff William Lichtman et al. in Cell. Published online June 1 2015 doi:10.1016/j.cell.2015.06.054
Abstract
Saturated Reconstruction of a Volume of Neocortex
Highlights
•Tape-based pipeline for electron microscopic reconstruction of brain tissue
•Annotated database of 1,700 synapses from a saturated reconstruction of cortex
•Excitatory axon proximity to dendritic spines not sufficient to predict synapses
Summary
We describe automated technologies to probe the structure of neural tissue at nanometer resolution and use them to generate a saturated reconstruction of a sub-volume of mouse neocortex in which all cellular objects (axons, dendrites, and glia) and many sub-cellular components (synapses, synaptic vesicles, spines, spine apparati, postsynaptic densities, and mitochondria) are rendered and itemized in a database. We explore these data to study physical properties of brain tissue. For example, by tracing the trajectories of all excitatory axons and noting their juxtapositions, both synaptic and non-synaptic, with every dendritic spine we refute the idea that physical proximity is sufficient to predict synaptic connectivity (the so-called Peters’ rule). This online minable database provides general access to the intrinsic complexity of the neocortex and enables further data-driven inquiries.
“Saturated Reconstruction of a Volume of Neocortex” by Jeff William Lichtman et al. in Cell. Published online June 1 2015 doi:10.1016/j.cell.2015.06.054