Summary: Researchers report they are able to see alterations in gene expression up to two hours after a stickleback fish interacts with an intruder. They speculate the changes in gene expression and chromatin modification could help set up the brain to respond to future threats. The study appears in PLOS Genetics.
Source: University of Illinois.
A five-minute encounter with an outsider spurs a cascade of changes in gene activity in the brain that can last for hours, researchers report in a study of stickleback fish.
The research, described in the journal PLOS Genetics, is one of three recent studies – the others conducted in honey bees and mice – to see waves of changes in gene expression in the brain 30 minutes to two hours after contact with an intruder.
“We are discovering that social interactions are extremely potent; they provoke big changes in gene expression in the brain,” said University of Illinois animal biology professor Alison Bell, who studies behavior in three-spined stickleback fish. “These very subtle social interactions are getting under the skin and becoming embedded in the brain. Studies like ours are beginning to show how that actually works.”
The observed changes included modifications to chromatin accessibility – a selective unpacking of the DNA that allows some genes, but not others, to be transcribed and translated into proteins.
“We used to think that DNA was so tightly packaged that it took a long time for that packaging to become unwrapped,” Bell said. “But our study shows that the unpackaging can happen fast – within minutes to hours – and we show that something really subtle, like a social interaction, can provoke dramatic changes in chromatin accessibility.”
Bell and her colleagues tracked changes in gene expression 30, 60 and 120 minutes after an encounter. They focused on two parts of the brain – the telencephalon, which is important to learning and memory; and the diencephalon, which integrates social information and hormonal influences.
“We compared experimental fish – those that had the encounter – with control fish, who remained alone in their tanks,” she said.
Hundreds of genes were expressed differently in the experimental and control fish. They also varied in the expression of numerous transcription factors, which regulate the activity of other genes.
Graduate Student Syed Abbas Bukhari, who is first author on the paper, led an analysis of the functions of groups of genes whose expression patterns paralleled one another at 30, 60 and 120 minutes.
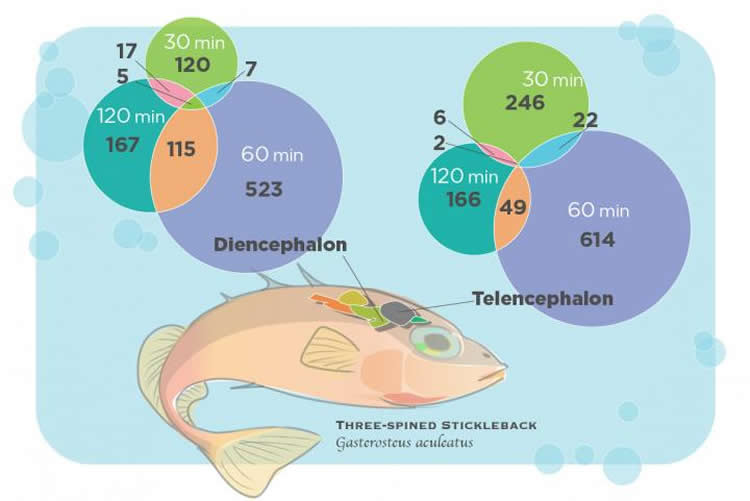
“This revealed that the genes that change in tandem over time have similar functions,” Bell said. Such functional groups include genes that contribute to immunity, hormone activity, metabolism or homeostasis.
In the experimental fish, hormone gene expression was highest at 30 minutes, metabolism genes peaked at 60 minutes and genes linked to immune function and homeostasis were upregulated the most two hours after the encounter.
“We’re seeing changes in gene expression even two hours after the fish interacts with an intruder. Clearly, those changes are not about the animal’s initial behavior,” Bell said. “We haven’t studied learning and memory, but we think that these chromatin modifications and changes in gene expression are all about setting up their brains to respond to threats in the future.”
Previous studies suggest that the fish are learning the limits of their territory, Bell said. “Their territory basically ends where they successfully chased away an intruder,” she said.
“Our findings made me feel very humble; I really did not appreciate how much is going on in these guys’ brains,” Bell said. “If you watch sticklebacks in natural populations, they’re constantly fighting with their neighbors. Our study just looked at what happened after a five-minute interaction, once. They’re doing this all day long, over and over again.”
Source: Diana Yates – University of Illinois
Image Source: NeuroscienceNews.com image is credited to Julie McMahon.
Original Research: Full open access research for “Temporal dynamics of neurogenomic plasticity in response to social interactions in male threespined sticklebacks” by Syed Abbas Bukhari, Michael C. Saul, Christopher H. Seward, Huimin Zhang, Miles Bensky, Noelle James, Sihai Dave Zhao, Sriram Chandrasekaran, Lisa Stubbs, and Alison M. Bell in PLOS Genetics. Published online July 13 2017 doi:10.1371/journal.pgen.1006840
[cbtabs][cbtab title=”MLA”]University of Illinois “Brief Interactions Spur Lasting Waves of Gene Activity in the Brain.” NeuroscienceNews. NeuroscienceNews, 17 July 2017.
<https://neurosciencenews.com/gene-activity-interactions-7104/>.[/cbtab][cbtab title=”APA”]University of Illinois (2017, July 17). Brief Interactions Spur Lasting Waves of Gene Activity in the Brain. NeuroscienceNew. Retrieved July 17, 2017 from https://neurosciencenews.com/gene-activity-interactions-7104/[/cbtab][cbtab title=”Chicago”]University of Illinois “Brief Interactions Spur Lasting Waves of Gene Activity in the Brain.” https://neurosciencenews.com/gene-activity-interactions-7104/ (accessed July 17, 2017).[/cbtab][/cbtabs]
Abstract
Temporal dynamics of neurogenomic plasticity in response to social interactions in male threespined sticklebacks
Animals exhibit dramatic immediate behavioral plasticity in response to social interactions, and brief social interactions can shape the future social landscape. However, the molecular mechanisms contributing to behavioral plasticity are unclear. Here, we show that the genome dynamically responds to social interactions with multiple waves of transcription associated with distinct molecular functions in the brain of male threespined sticklebacks, a species famous for its behavioral repertoire and evolution. Some biological functions (e.g., hormone activity) peaked soon after a brief territorial challenge and then declined, while others (e.g., immune response) peaked hours afterwards. We identify transcription factors that are predicted to coordinate waves of transcription associated with different components of behavioral plasticity. Next, using H3K27Ac as a marker of chromatin accessibility, we show that a brief territorial intrusion was sufficient to cause rapid and dramatic changes in the epigenome. Finally, we integrate the time course brain gene expression data with a transcriptional regulatory network, and link gene expression to changes in chromatin accessibility. This study reveals rapid and dramatic epigenomic plasticity in response to a brief, highly consequential social interaction.
“Temporal dynamics of neurogenomic plasticity in response to social interactions in male threespined sticklebacks” by Syed Abbas Bukhari, Michael C. Saul, Christopher H. Seward, Huimin Zhang, Miles Bensky, Noelle James, Sihai Dave Zhao, Sriram Chandrasekaran, Lisa Stubbs, and Alison M. Bell in PLOS Genetics. Published online July 13 2017 doi:10.1371/journal.pgen.1006840