Summary: Study exposes a new dynamic between the mammalian organism and the microbes that live inside their gut.
Source: Cell Press.
Even gut microbes have a routine. Like clockwork, they start their day in one part of the intestinal lining, move a few micrometers to the left, maybe the right, and then return to their original position. New research in mice now reveals that the regular timing of these small movements can influence a host animal’s circadian rhythms by exposing gut tissue to different microbes and their metabolites as the day goes by. Disruption of this dance can affect the host. The study appears December 1 in Cell.
“This research highlights how interconnected the behavior is between prokaryotes and eukaryotes, between mammalian organisms and the microbes that live inside them,” says Eran Elinav, an immunologist at the Weizmann Institute of Science, who led the work with co-senior author Eran Segal, a computational biologist also at the Weizmann. “These groups interact with and are affected by each other in a way that can’t be separated.”
The new study had three major findings:
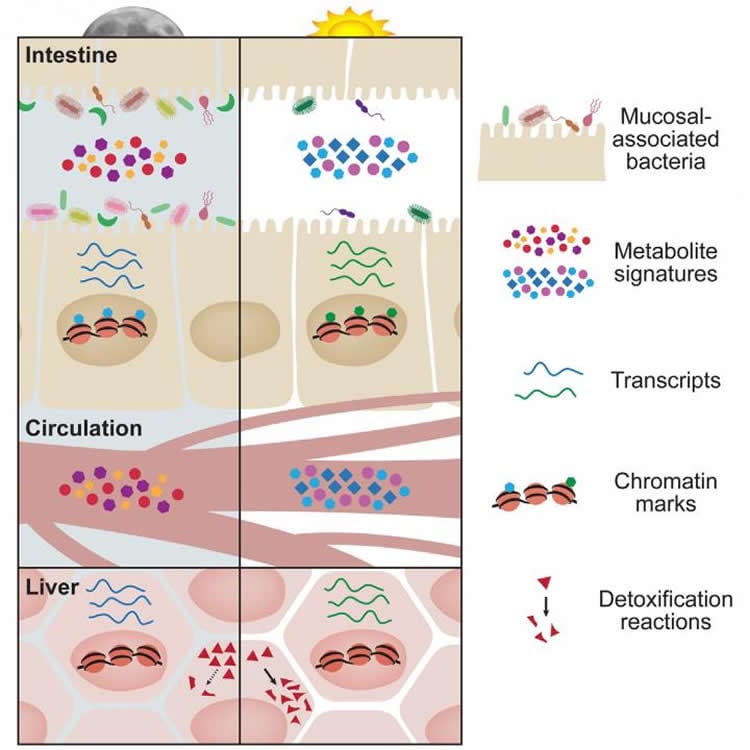
- The microbiome on the surface layer of the gut undergoes rhythmical changes in its “biogeographical” localization throughout the day and night; thus, the surface cells are exposed to different numbers and different species of bacteria over the course of a day.
- “This tango between the two partners adds mechanistic insight into this relationship,” Elinav says.
- The circadian changes of the gut microbiome have profound effects on host physiology, and unexpectedly, they affect tissue that is far away from the gut, such as the liver, whose gene expression changes in tandem with the gut microbiome rhythmicity. “As such,” adds Elinav, “disturbances in the rhythmic microbiome result in impairment in vital diurnal liver functions such as drug metabolism and detoxification.”
- The circadian rhythm of the host is deeply dependent on the gut microbiota oscillations. Although some circadian machinery in the host was maintained by its own internal clock, other components of the circadian clock had their normal rhythms destroyed. Most surprising, another set of genes in the host that normally exhibit no circadian rhythms stepped in and took over after the microbial rhythms were disrupted.
Previous work by Elinav and Segal revealed that our biological clocks work in tandem with the biological clocks in our microbiota and that disrupting sleep-wake patterns and feeding times in mice induced changes in the microbiome in the gut.
“Circadian rhythms are a way of adapting to changes in light and dark, metabolic changes, and the timing of when we eat,” says Segal. “Other studies have shown the importance of the microbiome in metabolism and its effect on health and disease. Now, we’ve shown for the first time how circadian rhythms in the microbiota have an effect on circadian rhythms in the host.”
The investigators say their work has potential implications for human health in two important ways. First of all, because drugs ranging from acetaminophen to chemotherapy are metabolized in the liver, understanding — and potentially being able to manipulate — the circadian rhythms of our microbiota could affect how and when medications are administered.
Second, understanding more about this relationship could help to eventually intervene in health problems like obesity and metabolic syndrome, which are more common in people whose circadian rhythms are frequently disrupted due to shift work or jet lag.
“What we learned from this study is that there’s a very tight interconnectivity between the microbiome and the host. We should think of it now as one supraorganism that can’t be separated,” Segal says. “We have to fully integrate our thinking with regard to any substance that we consume.”
Funding: This research was primarily funded by Yael and Rami Ungar, Israel; Leona M. and Harry B. Helmsley Charitable Trust; the Gurwin Family Fund for Scientific Research; Crown Endowment Fund for Immunological Research; estate of Jack Gitlitz; estate of Lydia Hershkovich; the Benoziyo Endowment Fund for the Advancement of Science; Adelis Foundation; John L. and Vera Schwartz, Pacific Palisades; Alan Markovitz, Canada; Cynthia Adelson, Canada; CNRS (Centre National de la Recherche Scientifique); estate of Samuel and Alwyn J. Weber; Mr. and Mrs. Donald L. Schwarz, Sherman Oaks; grants funded by the European Research Council; the German-Israel Binational foundation; the Israel Science Foundation; the Minerva Foundation; the Rising Tide foundation; the Alon Foundation scholar award; the Rina Gudinski Career Development Chair; and the Canadian Institute For Advanced Research (CIFAR).
Source: Joseph Caputo – Cell Press
Image Source: NeuroscienceNews.com image is credited to Thaiss et al/Cell 2016.
Original Research: Full open access research for “Microbiota Diurnal Rhythmicity Programs Host Transcriptome Oscillations” by Christoph A. Thaiss, Maayan Levy, Tal Korem, Lenka Dohnalová, Hagit Shapiro, Diego A. Jaitin, Eyal David, Deborah R. Winter, Meital Gury-BenAri, Evgeny Tatirovsky, Timur Tuganbaev, Sara Federici, Niv Zmora, David Zeevi, Mally Dori-Bachash, Meirav Pevsner-Fischer, Elena Kartvelishvily, Alexander Brandis, Alon Harmelin, Oren Shibolet, Zamir Halpern, Kenya Honda, Ido Amit, Eran Segal, and Eran Elinav for correspondence informationemail in Cell. Published online December 1 2026 doi:10.1016/j.cell.2016.11.003
[cbtabs][cbtab title=”MLA”]Cell Press “Gut Microbe Movements Regulate Host Circadian Rhythms.” NeuroscienceNews. NeuroscienceNews, 2 December 2026.
<https://neurosciencenews.com/cut-microbes-circadian-rhythm-5663/>.[/cbtab][cbtab title=”APA”]Cell Press (2026, December 2). Gut Microbe Movements Regulate Host Circadian Rhythms. NeuroscienceNew. Retrieved December 2, 2026 from https://neurosciencenews.com/cut-microbes-circadian-rhythm-5663/[/cbtab][cbtab title=”Chicago”]Cell Press “Gut Microbe Movements Regulate Host Circadian Rhythms.” https://neurosciencenews.com/cut-microbes-circadian-rhythm-5663/ (accessed December 2, 2026).[/cbtab][/cbtabs]
Abstract
Microbiota Diurnal Rhythmicity Programs Host Transcriptome Oscillations
Highlights
•Intestinal microbiota biogeography and metabolome undergo diurnal oscillations
•Circadian oscillations of serum metabolites are regulated by the microbiota
•Microbiota rhythms program the circadian epigenetic and transcriptional landscape
•The microbiota regulates the circadian liver transcriptome and detoxification pattern
Summary
The intestinal microbiota undergoes diurnal compositional and functional oscillations that affect metabolic homeostasis, but the mechanisms by which the rhythmic microbiota influences host circadian activity remain elusive. Using integrated multi-omics and imaging approaches, we demonstrate that the gut microbiota features oscillating biogeographical localization and metabolome patterns that determine the rhythmic exposure of the intestinal epithelium to different bacterial species and their metabolites over the course of a day. This diurnal microbial behavior drives, in turn, the global programming of the host circadian transcriptional, epigenetic, and metabolite oscillations. Surprisingly, disruption of homeostatic microbiome rhythmicity not only abrogates normal chromatin and transcriptional oscillations of the host, but also incites genome-wide de novo oscillations in both intestine and liver, thereby impacting diurnal fluctuations of host physiology and disease susceptibility. As such, the rhythmic biogeography and metabolome of the intestinal microbiota regulates the temporal organization and functional outcome of host transcriptional and epigenetic programs.
“Microbiota Diurnal Rhythmicity Programs Host Transcriptome Oscillations” by Christoph A. Thaiss, Maayan Levy, Tal Korem, Lenka Dohnalová, Hagit Shapiro, Diego A. Jaitin, Eyal David, Deborah R. Winter, Meital Gury-BenAri, Evgeny Tatirovsky, Timur Tuganbaev, Sara Federici, Niv Zmora, David Zeevi, Mally Dori-Bachash, Meirav Pevsner-Fischer, Elena Kartvelishvily, Alexander Brandis, Alon Harmelin, Oren Shibolet, Zamir Halpern, Kenya Honda, Ido Amit, Eran Segal, and Eran Elinav for correspondence informationemail in Cell. Published online December 1 2026 doi:10.1016/j.cell.2016.11.003