Summary: A new study reports on the benefits of precision medicine for treating pediatric brain cancer.
Source: Harvard.
Precision medicine — in which diagnosis and treatments are keyed to the genetic susceptibilities of individual cancers — can play a major role in treating children with brain tumors, suggests a study by investigators at Dana-Farber/Boston Children’s Cancer and Blood Disorders Center.
“Although there has been a great deal of progress over the past 30 years in improving survival rates for children with cancer, advances in pediatric brain cancer haven’t been as dramatic,” says co-lead author Pratiti Bandopadhayay of Dana-Farber/Boston Children’s, who is also an instructor in pediatrics at Harvard Medical School (HMS).
“In a recent study, brain tumors accounted for 25 percent of all pediatric deaths attributed to cancer. In addition, many of the current therapies can result in long-term difficulties in cognitive or physical functioning,” adds Bandopadhayay.
In the largest clinical study to date of genetic abnormalities in pediatric brain tumors, researchers performed clinical testing on more than 200 tumor samples and found that a majority had genetic irregularities that could influence how the disease was diagnosed and/or treated with approved drugs or agents being evaluated in clinical trials. The findings, reported online today by the journal Neuro-Oncology, demonstrate that testing pediatric brain tumor tissue for genetic abnormalities is clinically feasible and that in many cases the results can guide patients’ treatment.
Since emerging from research labs more than a decade ago, targeted therapies for cancer have significantly improved the treatment of certain types of leukemia, digestive system tumors, and breast cancer, among other malignancies. (Pathologists and cytogeneticists performed the testing in a federally approved clinical laboratory — certified under Clinical Laboratory Improvement Amendments as the only type of labs in the United States whose findings can guide patient treatment. Dana-Farber/Boston Children’s, the researchers noted, is one of the few centers in the country to regularly analyze the genetics of patients’ pediatric brain tumors.)
The researchers plumbed the genomes of 203 pediatric brain tumor samples, representing all major subtypes of the disease. They analyzed 117 of the samples with OncoPanel testing, a technology that sequences the exomes — the sections of DNA that hold the blueprints for making specific cell proteins — for irregularities in 300 cancer-related genes. They also studied 146 samples tested with OncoCopy, which examines how many copies of genes are missing or overabundant within the tumor cells. Sixty samples underwent both forms of testing, which allowed researchers to explore whether combining the two tests was more powerful than each alone.
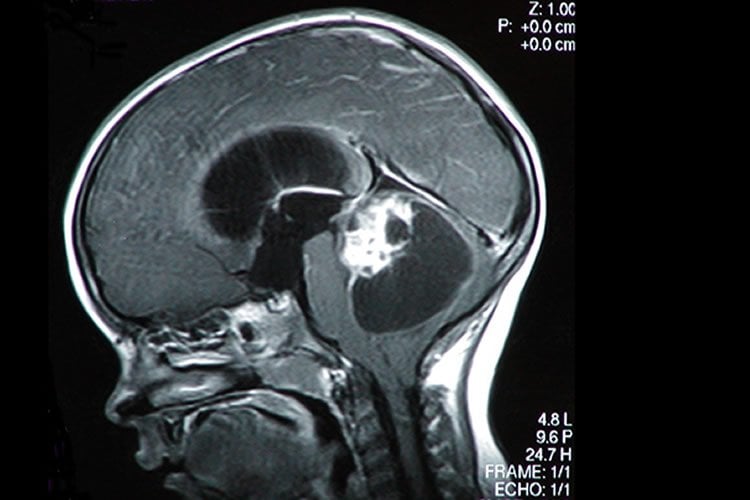
Of the samples tested by OncoPanel, 56 percent harbored genetic abnormalities that were clinically relevant —i.e., that could impact a patient’s diagnosis or be targeted by drugs already in clinical use or under study in clinical trials. (Many of these drugs cross the blood-brain barrier, the dense web of cells that can prevent medicines from exiting the bloodstream to reach the brain.)
Among the findings:
- Alterations were found in the gene BRAF, one of the most commonly mutated genes in pediatric brain tumors, and one for which several targeted drugs are being tested.
- The two-pronged testing approach revealed clinically relevant abnormalities in 89 percent of medulloblastomas, which account for nearly a fifth of all brain tumors in children. Combining the two tests was found to be particularly useful for these patients.
“The importance of genomic profiling in the diagnosis and treatment of pediatric brain cancers is reflected in the World Health Organization’s recent decision to classify such tumors by the genetic alterations within them, rather than by broad tumor type” says study co-senior author Susan Chi of Dana-Farber/Boston Children’s, an assistant professor of pediatrics at HMS. “Targeted therapies are likely to be most effective when they’re matched to specific abnormalities within tumor cells. Our findings show that precision medicine for pediatric brain tumors can now be a reality.”
The co-lead authors, with Bandopadhayay, of the study are: Shakti Ramkissoon of Dana-Farber Cancer Institute and Brigham and Women’s Hospital, Jaeho Hwang of Harvard Medical School, and Lori Ramkissoon of Dana-Farber. Co-senior authors, with Chi, are Rameen Beroukhim of Dana-Farber, Brigham and Women’s, and the Broad Institute of Harvard and MIT, and Keith Ligon of Dana-Farber/Boston Children’s, Brigham and Women’s, and the Broad Institute.
Source: Robert Levy – Harvard
Image Source: NeuroscienceNews.com image is credited to Google Images and is adapted from the Harvard press release.
Original Research: Abstract for “Clinical targeted exome-based sequencing in combination with genome-wide copy number profiling: precision medicine analysis of 203 pediatric brain tumors” by Ramkissoon, S. H., Bandopadhayay, P., Hwang, J., Ramkissoon, L. A., Greenwald, N. F., Schumacher, S. E., ORourke, R., Pinches, N., Ho, P., Malkin, H., Sinai, C., Filbin, M., Plant, A., Bi, W. L., Chang, M. S., Yang, E., Wright, K. D., Manley, P. E., Ducar, M., Alexandrescu, S., Lidov, H., Delalle, I., Goumnerova, L. C., Church, A. J., Janeway, K. A., Harris, M. H., MacConaill, L. E., Folkerth, R. D., Lindeman, N. I., Stiles, C. D., Kieran, M. W., Ligon, A. H., Santagata, S., Dubuc, A. M., Chi, S. N., Beroukhim, R., and Ligon, K. L. in Neuro-Oncology. Published online January 19 2017 doi:10.1093/neuonc/now294
[cbtabs][cbtab title=”MLA”]Harvard “New Hope For Children With Brain Tumors.” NeuroscienceNews. NeuroscienceNews, 21 January 2017.
<https://neurosciencenews.com/precision-medicine-brain-cancer-5985/>.[/cbtab][cbtab title=”APA”]Harvard (2017, January 21). New Hope For Children With Brain Tumors. NeuroscienceNew. Retrieved January 21, 2017 from https://neurosciencenews.com/precision-medicine-brain-cancer-5985/[/cbtab][cbtab title=”Chicago”]Harvard “New Hope For Children With Brain Tumors.” https://neurosciencenews.com/precision-medicine-brain-cancer-5985/ (accessed January 21, 2017).[/cbtab][/cbtabs]
Abstract
Clinical targeted exome-based sequencing in combination with genome-wide copy number profiling: precision medicine analysis of 203 pediatric brain tumorsn
Background. Clinical genomics platforms are needed to identify targetable alterations, but implementation of these technologies and best practices in routine clinical pediatric oncology practice are not yet well established.
Methods. Profile is an institution-wide prospective clinical research initiative that uses targeted sequencing to identify targetable alterations in tumors. OncoPanel, a multiplexed targeted exome-sequencing platform that includes 300 cancer-causing genes, was used to assess single nucleotide variants and rearrangements/indels. Alterations were annotated (Tiers 1–4) based on clinical significance, with Tier 1 alterations having well-established clinical utility. OncoCopy, a clinical genome-wide array comparative genomic hybridization (aCGH) assay, was also performed to evaluate copy number alterations and better define rearrangement breakpoints.
Results. Cancer genomes of 203 pediatric brain tumors were profiled across histological subtypes, including 117 samples analyzed by OncoPanel, 146 by OncoCopy, and 60 tumors subjected to both methodologies. OncoPanel revealed clinically relevant alterations in 56% of patients (44 cancer mutations and 20 rearrangements), including BRAF alterations that directed the use of targeted inhibitors. Rearrangements in MYB-QKI, MYBL1, BRAF, and FGFR1 were also detected. Furthermore, while copy number profiles differed across histologies, the combined use of OncoPanel and OncoCopy identified subgroup-specific alterations in 89% (17/19) of medulloblastomas.
Conclusion. The combination of OncoPanel and OncoCopy multiplex genomic assays can identify critical diagnostic, prognostic, and treatment-relevant alterations and represents an effective precision medicine approach for clinical evaluation of pediatric brain tumors.
“Clinical targeted exome-based sequencing in combination with genome-wide copy number profiling: precision medicine analysis of 203 pediatric brain tumors” by Ramkissoon, S. H., Bandopadhayay, P., Hwang, J., Ramkissoon, L. A., Greenwald, N. F., Schumacher, S. E., ORourke, R., Pinches, N., Ho, P., Malkin, H., Sinai, C., Filbin, M., Plant, A., Bi, W. L., Chang, M. S., Yang, E., Wright, K. D., Manley, P. E., Ducar, M., Alexandrescu, S., Lidov, H., Delalle, I., Goumnerova, L. C., Church, A. J., Janeway, K. A., Harris, M. H., MacConaill, L. E., Folkerth, R. D., Lindeman, N. I., Stiles, C. D., Kieran, M. W., Ligon, A. H., Santagata, S., Dubuc, A. M., Chi, S. N., Beroukhim, R., and Ligon, K. L. in Neuro-Oncology. Published online January 19 2017 doi:10.1093/neuonc/now294