Summary: Researchers propose future gut microbiome studies related to Parkinson’s disease should address questions relevant to patient care.
Source: IOS Press
Since the discovery that the gut microbiome may play a role in the development of Parkinson’s disease (PD), this fresh scientific approach has produced varying results. In this review published in the Journal of Parkinson’s Disease scientists compare results of current research and provide recommendations to increase the comparability and utility of these studies with a view towards improving patient outcomes.
“Despite the large societal impact of PD, the underlying cause remains elusive and only symptomatic treatments exist,” explained Filip Scheperjans, MD, PhD, Department of Neurology, Helsinki University Hospital, Helsinki, Finland, lead author of the review and chief investigator of the study that first identified gut microbiome differences in PD. “Since our study appeared in 2015, 15 human case-control studies with original data have found changes in the gut microbiome composition of PD patients, but their results vary. The challenge is to identify whether these changes are actually relevant for PD patients and whether they play a role in the disease process.”
“As more studies investigate the gut microbiome composition in PD, it is important to compare the findings of these studies to get an overview of the changes present in the disease,” added the review’s first author, Jeffrey M. Boertien, MSc, Parkinson Expertise Center, Department of Neurology, University Medical Center Groningen, University of Groningen, The Netherlands. “It is even more important to compare the methods used in the various studies. Especially as the studies report different and sometimes even contradictory results.”
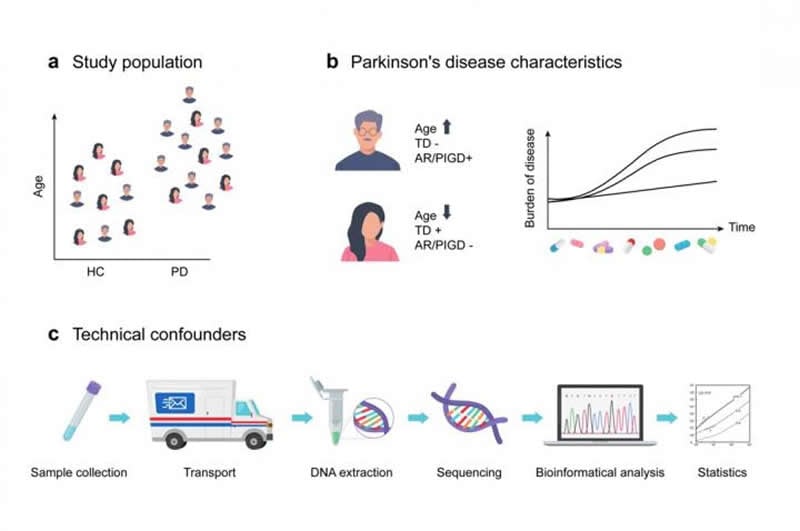
This review systematically compares the methodologies and results of the currently available case-control fecal gut microbiome studies in PD, which encompass 16 studies, representing seven countries with study populations varying from 10-197 PD patients and 10-130 healthy controls. These studies reported over 100 differentially abundant taxa covering all taxonomic levels, from phylum to genus or species, depending on methodology.
While several findings were replicated in various studies, such as an increase of Verrucomicrobiaceae and Akkermansia and a decrease of Prevotellaceae, the investigators also found numerous differences. “There is currently no consensus on PD-specific changes in microbiome composition and their pathophysiological implications due to inconsistent results, differences in methodologies and unaddressed confounders,” observed Dr. Scheperjans. Accordingly, procedures for collection, storage and shipment of the stool samples varied considerably; almost all studies used different DNA extraction kits; different DNA sequencing protocols were used; and different bioinformatics and statistical methods can further lead to different results. In addition, the study populations differed considerably between studies in terms of age, percentage of females and Parkinson’s disease characteristics, such as disease duration and the clinical subtype.
The investigators recommend methods to increase the comparability and utility of both the currently available data as well as future microbiome studies in PD. They propose the integration of the already available data to address various possible confounders. They also propose that future gut microbiome studies should address questions that are relevant for patient care, for example, whether gut microbiome changes can distinguish PD from similar disorders such as multiple system atrophy.
“Specific changes might serve as a biomarker with which we can recognize PD or specific subtypes of PD. Since gut complaints can occur very early in the disease process, this might help to identify patients in the early stages of the disease, possibly even before the appearance of motor symptoms such as tremor and rigidity,” commented Jeffrey Boertien. “If gut microbiota play an important role in the disease process, this might lead to new treatment options for PD.”
“If we combine all data, it will be easier to distinguish changes that are associated with PD from noise,” added Dr. Scheperjans. “However, further research is still required to increase our understanding of the possible role of gut microbiota in PD. It is important to emphasize that no microbiota-based treatment for PD exists to date. We advise PD patients not to start self-treatment with probiotics or undergo fecal microbiota transplantation without consulting with their doctors in order to avoid potential harm.”
PD is a slowly progressive disorder that affects movement, muscle control and balance. It is the second most common age-related neurodegenerative disorder affecting about 3% of the population by the age of 65 and up to 5% of individuals over 85 years of age. During the 20th century, PD was thought to be primarily a brain disorder. However, it is often preceded by non-motor symptoms such as sleep disorder, depression and gastrointestinal symptoms, especially constipation. The pathology present in the brains of PD patients, so-called Lewy bodies, can also be found in the nerve cells of the gut, leading to the hypothesis that PD might originate in the gut.
Source:
IOS Press
Media Contacts:
Diana Murray – IOS Press
Image Source:
The image is credited to Academy of Finland.
Original Research: Open access
“Increasing Comparability and Utility of Gut Microbiome Studies in Parkinson’s Disease: A Systematic Review”. Filip Scheperjans et al.
Journal of Parkinson’s Disease doi:10.3233/JPD-191711.
Abstract
Increasing Comparability and Utility of Gut Microbiome Studies in Parkinson’s Disease: A Systematic Review
Gut microbiota have been studied in relation to the pathophysiology of Parkinson’s disease (PD) due to the early gastrointestinal symptomatology and presence of alpha-synuclein pathology in the enteric nervous system, hypothesized to ascend via the vagal nerve to the central nervous system. Accordingly, sixteen human case-control studies have published gut microbiome composition changes in PD and reported over 100 differentially abundant taxa covering all taxonomic levels from phylum to genus or species, depending on methodology. While certain findings were replicated across several studies, various contradictory findings were reported. Here, differences in methodologies and the presence of possible confounders in the study populations are assessed for their potential to confound the results of gut microbiome studies in PD. Gut microbiome studies in PD exhibited considerable variability with respect to the study population, sample transport conditions, laboratory protocols and sequencing, bioinformatics pipelines, and biostatistical methods. To move from the current heterogeneous dataset towards clinically relevant biomarkers and the identification of putative therapeutic targets, recommendations are derived from the limitations of the available studies to increase the future comparability of microbiome studies in PD. In addition, integration of currently available data on the gut microbiome in PD is proposed to identify robust gut microbiome profiles in PD. Furthermore, expansion of the current dataset with atypical parkinsonism cohorts, prodromal and treatment-naïve de novo PD subjects, measurements of fecal microbial concentrations and multi-omics assessments are required to provide clinically relevant biomarkers and reveal therapeutic targets within the gut microbiome of PD.