Summary: CRISPR technology and advanced screening techniques allow researchers to comb through over 1500 genetic combinations to find multiple drivers of glioblastoma brain cancer.
Source: Yale.
From thousands of suspects, Yale scientists ferret out cancer-causing genes.
A Yale-led team of researchers has identified specific gene combinations that can cause the deadly brain cancer glioblastoma, using new technology that can also pinpoint triggers of other types cancers, they report Aug. 14 in the journal Nature Neuroscience.
Scientists have been adept at identifying mutations present in a variety of cancers in patients, but it is still challenging for researchers to identify the genes or combinations of genes that are directly causing the progression of the disease. For instance, more than 223 individual genes have been linked to glioblastoma, a difficult-to-treat brain cancer with a median survival rate of only 1 to 1.5 years. Since thousands of combinations of those genes could act in concert to cause the disease in an individual patient, it has been challenging to determine which mutations are the most relevant for the growth of a given cancer.
“The human cancer genome is now mapped and thousands of new mutations were associated with cancer, but it has been difficult to prove which ones or their combinations actually cause cancer,” said Sidi Chen, assistant professor of genetics and at Yale’s Systems Biology Institute, and a co-corresponding author of the paper. “We can also use this information to determine which existing drugs are most likely to have therapeutic value for individual patients, a step towards personalized cancer therapy.’’
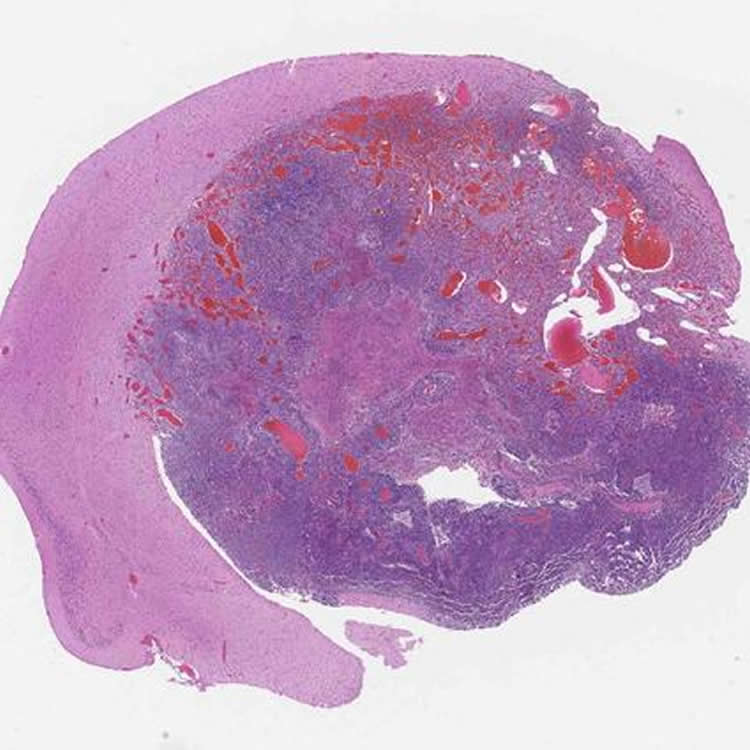
Chen, a member of the Yale Cancer Center, and colleagues developed an improved application of CRISPR gene editing and screening technology to search for primary drivers of glioblastoma in living mice. They assessed the impact of mutations in more than 1500 genetic combinations and found multiple combinations that could cause the cancer. They also found two mutations that could make tumors resistant to chemotherapy — information that could help doctors tailor existing treatments for individual patients, said the researchers.
The new technology could also provide researchers with ways to find specific targets for new drugs, Chen said.
Randall J. Platt at ETH Zurich is a co-corresponding author. Yale’s Ryan D Chow, Christopher D Guzman, Guangchuan Wang and ETH’s Florian Schmidt are co-first authors of the paper.
Funding: Primary funding for the research was provided by Yale System Biology Institute and the National Institutes of Health.
Source: Bill Hathaway – Yale
Image Source: NeuroscienceNews.com image is credited to Chen Lab.
Original Research: Abstract for “AAV-mediated direct in vivo CRISPR screen identifies functional suppressors in glioblastoma” by Ryan D Chow, Christopher D Guzman, Guangchuan Wang, Florian Schmidt, Mark W Youngblood, Lupeng Ye, Youssef Errami, Matthew B Dong, Michael A Martinez, Sensen Zhang, Paul Renauer, Kaya Bilguvar, Murat Gunel, Phillip A Sharp, Feng Zhang, Randall J Platt & Sidi Chen in Nature Neuroscience. Published online August 14 2017 doi:10.1038/nn.4620
[cbtabs][cbtab title=”MLA”]Yale “Gene Combinations That Cause Glioblastoma Brain Cancer Identified.” NeuroscienceNews. NeuroscienceNews, 14 August 2017.
<https://neurosciencenews.com/glioblastoma-gene-combinations-7298/>.[/cbtab][cbtab title=”APA”]Yale (2017, August 14). Gene Combinations That Cause Glioblastoma Brain Cancer Identified. NeuroscienceNew. Retrieved August 14, 2017 from https://neurosciencenews.com/glioblastoma-gene-combinations-7298/[/cbtab][cbtab title=”Chicago”]Yale “Gene Combinations That Cause Glioblastoma Brain Cancer Identified.” https://neurosciencenews.com/glioblastoma-gene-combinations-7298/ (accessed August 14, 2017).[/cbtab][/cbtabs]
Abstract
AAV-mediated direct in vivo CRISPR screen identifies functional suppressors in glioblastoma
A causative understanding of genetic factors that regulate glioblastoma pathogenesis is of central importance. Here we developed an adeno-associated virus–mediated, autochthonous genetic CRISPR screen in glioblastoma. Stereotaxic delivery of a virus library targeting genes commonly mutated in human cancers into the brains of conditional-Cas9 mice resulted in tumors that recapitulate human glioblastoma. Capture sequencing revealed diverse mutational profiles across tumors. The mutation frequencies in mice correlated with those in two independent patient cohorts. Co-mutation analysis identified co-occurring driver combinations such as B2m–Nf1, Mll3–Nf1 and Zc3h13–Rb1, which were subsequently validated using AAV minipools. Distinct from Nf1-mutant tumors, Rb1-mutant tumors are undifferentiated and aberrantly express homeobox gene clusters. The addition of Zc3h13 or Pten mutations altered the gene expression profiles of Rb1 mutants, rendering them more resistant to temozolomide. Our study provides a functional landscape of gliomagenesis suppressors in vivo.
“AAV-mediated direct in vivo CRISPR screen identifies functional suppressors in glioblastoma” by Ryan D Chow, Christopher D Guzman, Guangchuan Wang, Florian Schmidt, Mark W Youngblood, Lupeng Ye, Youssef Errami, Matthew B Dong, Michael A Martinez, Sensen Zhang, Paul Renauer, Kaya Bilguvar, Murat Gunel, Phillip A Sharp, Feng Zhang, Randall J Platt & Sidi Chen in Nature Neuroscience. Published online August 14 2017 doi:10.1038/nn.4620