Summary: Examining postmortem brains, researchers discover the state of glia remains consistent as we age and could be used to predict a person’s age at death.
Source: Cell Press.
The difference between an old brain and a young brain isn’t so much the number of neurons but the presence and function of supporting cells called glia. In Cell Reports on January 10, researchers who examined postmortem brain samples from 480 individuals ranging in age from 16 to 106 found that the state of someone’s glia is so consistent through the years that it can be used to predict someone’s age. The work lays the foundation to better understand glia’s role in late-in-life brain disease.
“We extensively characterized aging-altered gene expression changes across 10 human brain regions and found that, in fact, glial cells experience bigger changes than neurons,” says Jernej Ule, a neurobiologist at the Francis Crick Institute and the University College London, who led the study with departmental colleague Rickie Patani and first author Lilach Soreq. “There’s quite a bit of regional information that will be of interest to different people–for example some will notice a very unique pattern of astrocyte-specific changes in the substantia nigra–and we provide a lot of data that still needs to be analyzed.”
There are three types of glia cells, each providing different kinds of support to neurons: oligodendrocytes insulate, microglia act as immune cells, and astrocytes help with neuron metabolism, detoxification, among many functions. Based on analysis of human brain tissue samples, primarily from the UK Brain Expression Consortium, the researchers show that astrocytes and oligodendrocytes shift their regional gene expression patterns upon aging, (e.g., which genes are turned on or off) particularly in the hippocampus and substantia nigra–important brain regions for memory and movement, respectively–while the expression of microglia-specific genes increases in all brain regions.
The investigators next took a preliminary look at whether these changes in gene expression could relate to changes in brain cell populations. Based on a comparison of tissue samples from 3 young and 3 old brains, they found that the number of oligodendrocytes decreases with age in the frontal cortex. They further established that this likely corresponds with decreased expression of oligodendrocyte specific genes. Other types of cells had more complicated patterns of change.
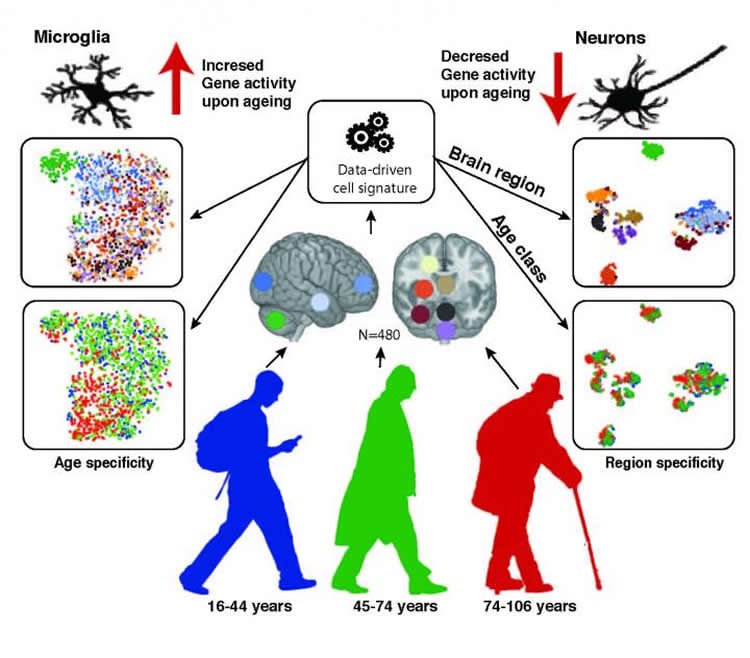
“We developed a very nice machine learning program and had to go through hundreds of thousands of oligodendrocytes and neurons to get reliable data, but we wanted to understand whether decreased expression causes changes at the molecular or cellular-level,” Ule says. “We did see oligodendrocytes disappearing but with neurons we didn’t see dramatic changes in cellular numbers except for a decrease in the largest neurons. This is of interest because those largest neurons are generally connected to neurodegenerative diseases.”
One unexpected finding was that certain glial gene expression patterns could predict age in the general population. While this can only be done postmortem, and certain people will not fit neatly into these patterns, it does provide scientists one more tool to understand how aging in the brain may be linked to the causes of age-related disorders. The researchers’ ultimate goal is to see whether gene mutations or other variables could affect gene expression in ways that cause disease.
Funding: This work was supported by the European Research Council; the Marie Curie Intra European Fellowship and Alzheimer’s Society for Junior Investigator award, the Francis Crick Institute, the UK Medical Research Council, the Wellcome Trust; the UK Medical Research Council; and the US National Institute on Aging, NIH.
Source: Joseph Caputo – Cell Press
Image Source: NeuroscienceNews.com image is credited to Lilach Soreq.
Original Research: Full open access research for “Major Shifts in Glial Regional Identity Are a Transcriptional Hallmark of Human Brain Aging” by Lilach Soreq, UK Brain Expression Consortium, North American Brain Expression Consortium, Jamie Rose, Eyal Soreq, John Hardy, Daniah Trabzuni, Mark R. Cookson, Colin Smith, Mina Ryten, Rickie Patani, and Jernej Ule in Cell Reports. Published online January 10 2017 doi:10.1016/j.celrep.2016.12.011
[cbtabs][cbtab title=”MLA”]Cell Press “Glia, Not Neurons, Most Affected By Brain Aging.” NeuroscienceNews. NeuroscienceNews, 10 January 2017.
<https://neurosciencenews.com/glia-brain-aging-5902/>.[/cbtab][cbtab title=”APA”]Cell Press (2017, January 10). Glia, Not Neurons, Most Affected By Brain Aging. NeuroscienceNew. Retrieved January 10, 2017 from https://neurosciencenews.com/glia-brain-aging-5902/[/cbtab][cbtab title=”Chicago”]Cell Press “Glia, Not Neurons, Most Affected By Brain Aging.” https://neurosciencenews.com/glia-brain-aging-5902/ (accessed January 10, 2017).[/cbtab][/cbtabs]
Abstract
Major Shifts in Glial Regional Identity Are a Transcriptional Hallmark of Human Brain Aging
Highlights
•Understanding the role of cell-type-specific changes in human brain aging
•Glial-specific genes shift their regional expression patterns during aging
•Oligodendrocytes and neuronal subpopulations are decreased in the aging neocortex
•Microglia-specific genes globally increase their expression during aging
Summary
Gene expression studies suggest that aging of the human brain is determined by a complex interplay of molecular events, although both its region- and cell-type-specific consequences remain poorly understood. Here, we extensively characterized aging-altered gene expression changes across ten human brain regions from 480 individuals ranging in age from 16 to 106 years. We show that astrocyte- and oligodendrocyte-specific genes, but not neuron-specific genes, shift their regional expression patterns upon aging, particularly in the hippocampus and substantia nigra, while the expression of microglia- and endothelial-specific genes increase in all brain regions. In line with these changes, high-resolution immunohistochemistry demonstrated decreased numbers of oligodendrocytes and of neuronal subpopulations in the aging brain cortex. Finally, glial-specific genes predict age with greater precision than neuron-specific genes, thus highlighting the need for greater mechanistic understanding of neuron-glia interactions in aging and late-life diseases.
“Major Shifts in Glial Regional Identity Are a Transcriptional Hallmark of Human Brain Aging” by Lilach Soreq, UK Brain Expression Consortium, North American Brain Expression Consortium, Jamie Rose, Eyal Soreq, John Hardy, Daniah Trabzuni, Mark R. Cookson, Colin Smith, Mina Ryten, Rickie Patani, and Jernej Ule in Cell Reports. Published online January 10 2017 doi:10.1016/j.celrep.2016.12.011