Summary: Researchers have identified a set of genes that may have influenced human brain size, which helped us diverge from other apes. The study reveals the genes are unique to humans and help to build the neocortex.
Source: HHMI.
A newly discovered set of genes could help explain how humans diverged from other apes.
Three genes involved in nerve cell generation in the brain emerged roughly 3.5 million years ago and may have contributed to the rapid evolution of the large human brain, Howard Hughes Medical Institute (HHMI) Investigator David Haussler and colleagues report on May 31, 2018, in the journal Cell. That work – and a paper from an independent research team published in the same issue – identifies genes that help build the neocortex, the wrinkly outer layer of the brain that gives humans our ability to think, plan, and reason.
Until now, the genes had been unexplored, masked by an error in the published version of the human genome. These genes, and others unique to humans, offer clues about what separates us from chimpanzees, says Haussler, of the University of California, Santa Cruz (UCSC).
“Understanding ourselves and how we’re different from other species has been a scientific quest for millennia,” he says. “These studies help address the age-old question of what makes us human.”
Ancient history
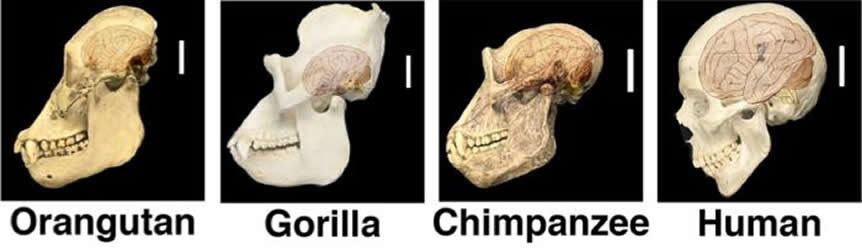
About 6.5 million years ago, humans and chimpanzees diverged from a common ancestor. A few million years after that, human brain size began to grow. “If you look at craniums from fossils of our ancestors, you can see this expansion,” Haussler says.
Today, the human neocortex is about three times larger than that of a chimp, but researchers haven’t identified all the genetic factors responsible for the difference. Since humans and chimps split, the human genome has undergone roughly 15 million changes – tweaks to the DNA letters that make up our genetic instruction book.
Haussler is well-versed in the particulars of people’s genes. His team was part of the Human Genome Project, the multiyear quest to spell out all of the DNA letters that make up humans’ roughly 20,000 genes. Haussler and colleagues posted the first draft of the human genome on the Internet on July 7, 2000. An official version came out in 2003, but it’s still a work in progress, he says. “We keep filling in little gaps and clarifying places that we were uncertain about.”
Researchers also have to decide which variants of genes to include. Humans all have the same basic blueprints, but genes can be rearranged, deleted, and come in different “flavors.” The official genome is “mostly some anonymous guy from Buffalo,” Haussler says, but scientists have swapped out some of his altered genes to build a more complete version of the genome.
They’re now on version 38, and each new iteration can unearth valuable nuggets of information. For Haussler’s team, version 38 struck gold.
Missing pieces
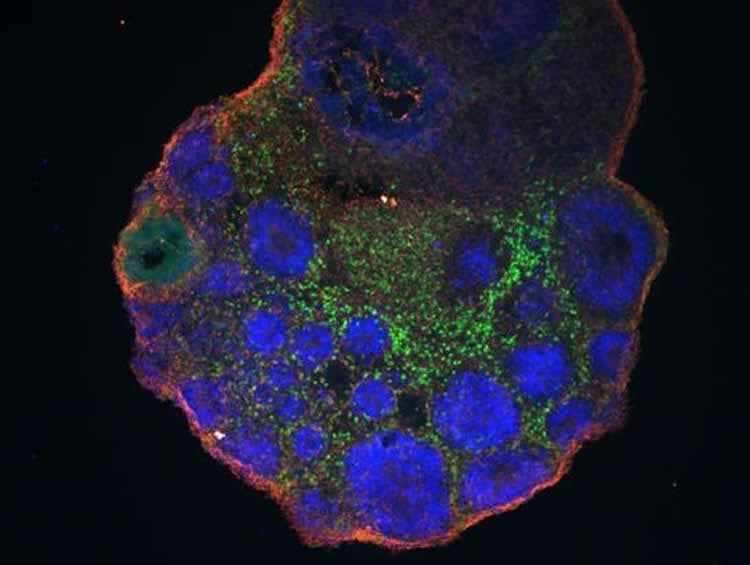
Haussler and his colleagues had been growing embryonic brain-like structures in the lab and looking for differences between those made from humans and rhesus monkeys. His team was trying to pinpoint genes that were active only in developing human brains. Such genes could include a key to human brain size - and human intelligence.
About six years ago, study coauthor Frank Jacobs, then in Haussler’s lab and now at the University of Amsterdam, discovered a promising candidate – a gene called NOTCH2NL. It’s a relative of NOTCH2, a gene that scientists knew was central to early brain development, Haussler says. NOTCH2 controls vital decisions regarding when and how many neurons to make.
When the Haussler team looked in the official version of the human genome at that time – version 37 – NOTCH2NL appeared to be located in chromosome 1 near a region linked to abnormal brain size. Delete a hunk of the region, and brains tend to shrink. Duplicate part of it, and brains tend to overgrow.
“We thought, ‘Oh, this is incredible,'” Haussler says. NOTCH2NL seemed to check all the boxes for a key role in human brain development. But when the team mapped NOTCH2NL’s precise location in the genome, they discovered the gene wasn’t actually in the relevant chromosomal region after all; the once-promising candidate seemed to be a dud.
“We were downhearted,” Haussler says. That all changed with the next official version of the human genome – version 38. In this iteration, NOTCH2NL was located in the crucial region – “And there were three versions of it!” he says. Over the last three million years, his team calculated, NOTCH2NL was repeatedly copy-pasted into the genome, what he calls “a series of genetic accidents.”
Genetic analysis of several primate species revealed that the three genes exist only in humans and their recent relatives, the Neanderthals and Denisovans, not in chimpanzees, gorillas, or orangutans. What’s more, the timing of these genes’ emergence matches up with the period in the fossil record when our ancestors’ craniums began to enlarge, Haussler says. Together, the results suggest that NOTCH2NL genes played a role in beefing up human brain size.
“I thought, oh my God, now it all really fits together,” he says.
Brain boosters
UCSC cell biologist Sofie Salama led efforts to figure out how these genes might work – and how they might contribute to disease.
Experiments with lab-grown patches of brain tissue from mouse and human embryonic stem cells hinted that NOTCH2NL genes tell developing brains to make more neuron progenitors. These are like little neuron factories, Haussler says. More progenitors mean more neurons – and ultimately bigger brains.
The NOTCH2NL genes likely convey this brain-growing message via the well-studied Notch signaling pathway, Salama and her colleagues discovered. Scientists already knew that cells rely on this pathway for proper neuronal function. Having three active NOTCH2NL genes cranks up the messaging, “adding an extra push to the pathway,” Salama says.
Deviations in these genes’ copy numbers could be tied to neurological disorders other than abnormal brain size, the researchers found. An analysis of cells from six autism patients in the Simons Variation in Individuals project revealed that each patient had lost or gained parts of NOTCH2NL genes. Until now, scientists hadn’t mapped the exact genomic regions duplicated or cropped out.
Precise mapping was difficult, Salama says, because that area of the genome is so repetitive – assembling the correct sequence of DNA is like trying to put together a puzzle with nearly identical pieces.
But the team developed a sequencing technology to solve the problem, and pinpointed how the NOTCH2NL genes were improperly pieced together in the patients with autism. The results suggest a direct link between the disorder and reshuffled NOTCH2NL genes, Salama says.
Haussler’s team’s findings jibe with those from the University of Brussels (ULB) lab of neuroscientist Pierre Vanderhaeghen, whose paper appears alongside Haussler’s in Cell. Vanderhaeghen and his colleagues designed a computational method to help detect genes active during human fetal brain development. The top gene on the list was the same one Haussler identified – NOTCH2NL. “It’s very exciting that we picked up the same super-cool candidate,” Vanderhaeghen says.
The work showcases the importance of gene duplication as a force in evolution, says HHMI Investigator Christopher Walsh, of Boston Children’s Hospital. Multiple copies of the same gene can drive major changes in the body. Walsh recently reported that a single copy of a different gene, ASPM, could have also played a role in human brain evolution.
These genes and others like them are already spurring new questions. In modern humans, for example, NOTCH2NL exists in at least eight slightly different iterations – subtle variations in the order of DNA letters that make up the gene. Haussler says it’s unclear what effect these differences may have. And Vanderhaeghen’s team has other candidate neurodevelopmental genes on the list that includes NOTCH2NL. “These are the top candidates of human-specific genes,” he says. “No one has studied them yet.”
A deep dive into the genome could be a gateway to explaining human evolution and brainpower, Haussler says. “Understanding the key to our greater cognitive abilities is a huge opportunity of our time. This is an area for young scientists to be excited about.”
Funding: Funding provided by Natural Science and Engineering Research Council of Canada, Canadian Institutes of Health Research.
Source: Meghan Rosen – HHMI
Publisher: Organized by NeuroscienceNews.com.
Image Source: NeuroscienceNews.com images are credited to Vincent Meng and Andrew Field, and Richard Baldwin of the Anthropology Department at UCSC.
Original Research: Open access research for “Human-Specific NOTCH2NL Genes Affect Notch Signaling and Cortical Neurogenesis” by Ian T. Fiddes, Gerrald A. Lodewijk, Meghan Mooring, Colleen M. Bosworth, Adam D. Ewing, Gary L. Mantalas, Adam M. Novak, Anouk van den Bout, Alex Bishara, Jimi L. Rosenkrantz, Ryan Lorig-Roach, Andrew R. Field, Maximilian Haeussler, Lotte Russo, Aparna Bhaduri, Tomasz J. Nowakowski, Alex A. Pollen, Max L. Dougherty, Xander Nuttle, Marie-Claude Addor, Simon Zwolinski, Sol Katzman, Arnold Kriegstein, Evan E. Eichler, Sofie R. Salama14, Frank M.J. Jacobs, David Haussler in Cell. Published May 31 2018.
doi:10.1016/j.cell.2018.03.051
[cbtabs][cbtab title=”MLA”]HHMI “Three New Genes That May Have Influenced Human Brain Size.” NeuroscienceNews. NeuroscienceNews, 31 May 2018.
<https://neurosciencenews.com/genetics-brain-size-9202/>.[/cbtab][cbtab title=”APA”]HHMI (2018, May 31). Three New Genes That May Have Influenced Human Brain Size. NeuroscienceNews. Retrieved May 31, 2018 from https://neurosciencenews.com/genetics-brain-size-9202/[/cbtab][cbtab title=”Chicago”]HHMI “Three New Genes That May Have Influenced Human Brain Size.” https://neurosciencenews.com/genetics-brain-size-9202/ (accessed May 31, 2018).[/cbtab][/cbtabs]
Abstract
Human-Specific NOTCH2NL Genes Affect Notch Signaling and Cortical Neurogenesis
Highlights
•NOTCH2NLA,B,C expressed in human fetal brain radial glia stem cells, arose 0.5–4 mya
•These genes encode secreted, Notch-related proteins that enhance Notch signaling
•Overexpression of NOTCH2NL delays neuronal differentiation, while deletion accelerates it
•NOTCH2NLA and NOTCH2NLB serve as breakpoints in 1q21.1 deletion/duplication syndrome
Summary
Genetic changes causing brain size expansion in human evolution have remained elusive. Notch signaling is essential for radial glia stem cell proliferation and is a determinant of neuronal number in the mammalian cortex. We find that three paralogs of human-specific NOTCH2NL are highly expressed in radial glia. Functional analysis reveals that different alleles of NOTCH2NL have varying potencies to enhance Notch signaling by interacting directly with NOTCH receptors. Consistent with a role in Notch signaling, NOTCH2NL ectopic expression delays differentiation of neuronal progenitors, while deletion accelerates differentiation into cortical neurons. Furthermore, NOTCH2NL genes provide the breakpoints in 1q21.1 distal deletion/duplication syndrome, where duplications are associated with macrocephaly and autism and deletions with microcephaly and schizophrenia. Thus, the emergence of human-specific NOTCH2NL genes may have contributed to the rapid evolution of the larger human neocortex, accompanied by loss of genomic stability at the 1q21.1 locus and resulting recurrent neurodevelopmental disorders.