Summary: Using a highly versatile form of CRISPR gene editing, researchers successfully restored vision in mice with retinitis pigmentosa.
Source: Rockefeller University Press
Researchers in China have successfully restored the vision of mice with retinitis pigmentosa, one of the major causes of blindness in humans.
The study, to be published March 17 in the Journal of Experimental Medicine, uses a new, highly versatile form of CRISPR-based genome editing with the potential to correct a wide variety of disease-causing genetic mutations.
Researchers have previously used genome editing to restore the vision of mice with genetic diseases, such as Leber congenital amaurosis, that affect the retinal pigment epithelium, a layer of non-neuronal cells in the eye that supports the light-sensing rod and cone photoreceptor cells. However, most inherited forms of blindness, including retinitis pigmentosa, are caused by genetic defects in the neural photoreceptors themselves.
“The ability to edit the genome of neural retinal cells, particularly unhealthy or dying photoreceptors, would provide much more convincing evidence for the potential applications of these genome-editing tools in treating diseases such as retinitis pigmentosa,” says Kai Yao, a professor at the Wuhan University of Science and Technology.
Retinitis pigmentosa can be caused by mutations in over 100 different genes and is estimated to impair the vision of 1 in 4,000 people. It begins with the dysfunction and death of dim light-sensing rod cells, before spreading to the cone cells required for color vision, eventually leading to severe, irreversible vision loss.
Yao and colleagues attempted to rescue the vision of mice with retinitis pigmentosa caused by a mutation in the gene encoding a critical enzyme called PDE6β. To do this, Yao’s team developed a new, more versatile CRISPR system called PESpRY, which can be programmed to correct many different types of genetic mutation, regardless of where they occur within the genome.
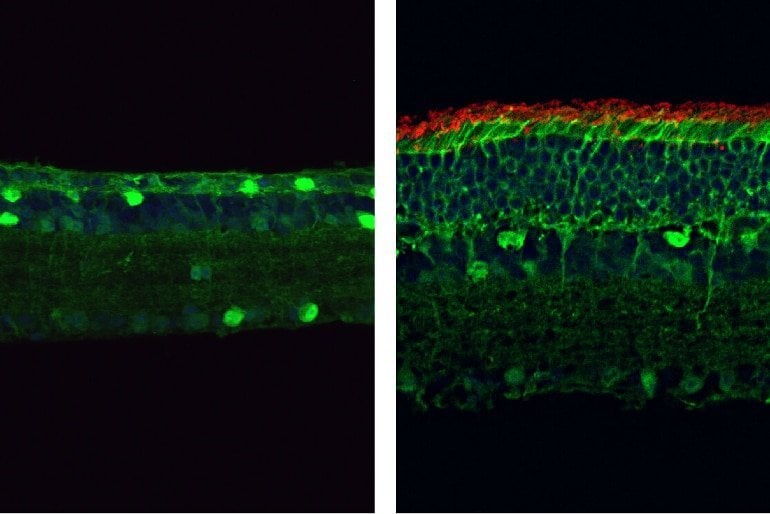
When programmed to target the mutant PDE6β gene, the PESpRY system was able to efficiently correct the mutation and restore the enzyme’s activity in the retinas of mice. This prevented the death of rod and cone photoreceptors and restored their normal electrical responses to light.
Yao and colleagues performed a variety of behavioral tests to confirm that the gene-edited mice retained their vision even into old age. For example, the animals were able to find their way out of a visually guided water maze almost as well as normal, healthy mice and showed typical head movements in response to visual stimuli.
Yao cautions that much work still needs to be done to establish both the safety and efficacy of the PESpRY system in humans.
“However, our study provides substantial evidence for the in vivo applicability of this new genome-editing strategy and its potential in diverse research and therapeutic contexts, in particular for inherited retinal diseases such as retinitis pigmentosa,” Yao says.
About this gene editing and visual neuroscience research news
Author: Press Office
Source: Rockefeller University Press
Contact: Press Office – Rockefeller University Press
Image: The image is credited to Qin et al / JEM
Original Research: Open access.
“Vision rescue via unconstrained in vivo prime editing in degenerating neural retinas” by Huan Qin et al. Journal of Experimental Medicine
Abstract
Vision rescue via unconstrained in vivo prime editing in degenerating neural retinas
Retinitis pigmentosa (RP) is an inherited retinal dystrophy causing progressive and irreversible loss of retinal photoreceptors.
Here, we developed a genome-editing tool characterized by the versatility of prime editors (PEs) and unconstrained PAM requirement of a SpCas9 variant (SpRY), referred to as PESpRY.
The diseased retinas of Pde6b-associated RP mouse model were transduced via a dual AAV system packaging PESpRY for the in vivo genome editing through a non-NGG PAM (GTG).
The progressing cell loss was reversed once the mutation was corrected, leading to substantial rescue of photoreceptors and production of functional PDE6β. The treated mice exhibited significant responses in electroretinogram and displayed good performance in both passive and active avoidance tests.
Moreover, they presented an apparent improvement in visual stimuli-driven optomotor responses and efficiently completed visually guided water-maze tasks.
Together, our study provides convincing evidence for the prevention of vision loss caused by RP-associated gene mutations via unconstrained in vivo prime editing in the degenerating retinas.







