Summary: A new study reports microglia may play a role in a diverse array of neurodegenerative and psychiatric illnesses.
Source: Salk Institute.
Salk and UC San Diego scientists conducted vast microglia survey, revealing links to neurodegenerative diseases and psychiatric illnesses.
Scientists have, for the first time, characterized the molecular markers that make the brain’s front lines of immune defense—cells called microglia—unique. In the process, they discovered further evidence that microglia may play roles in a variety of neurodegenerative and psychiatric illnesses, including Alzheimer’s, Parkinson’s and Huntington’s diseases as well as schizophrenia, autism and depression.
“Microglia are the immune cells of the brain, but how they function in the human brain is not well understood,” says Rusty Gage, professor in Salk’s Laboratory of Genetics, the Vi and John Adler Chair for Research on Age-Related Neurodegenerative Disease, and a senior author of the new work. “Our work not only provides links to diseases but offers a jumping off point to better understand the basic biology of these cells.”
Genes that have previously been linked to neurological diseases are turned on at higher levels in microglia compared to other brain cells, the team reported in Science on May 25, 2017. While the link between microglia and a number of disorders has been explored in the past, the new study offers a molecular basis for this connection.
“These studies represent the first systematic effort to molecularly decode microglia,” says Christopher Glass, a Professor of Cellular and Molecular Medicine and Professor of Medicine at University of California San Diego, also senior author of the paper. “Our findings provide the foundations for understanding the underlying mechanisms that determine beneficial or pathological functions of these cells.”
Microglia are a type of macrophage, white blood cells found throughout the body that can destroy pathogens or other foreign materials. They’re known to be highly responsive to their surroundings and respond to changes in the brain by releasing pro-inflammatory or anti-inflammatory signals. They also prune back the connections between neurons when cells are damaged or diseased. But microglia are notoriously hard to study. They can’t be easily grown in a culture dish and quickly die outside of a living brain.
Nicole Coufal, a pediatric critical care doctor at UC San Diego, who also works in the Gage lab at Salk, wanted to make microglia from stem cells. But she realized there wasn’t any way to identify whether the resulting cells were truly microglia.
“There was not a unique marker that differentiated microglia from circulating macrophages in the rest of the body,” she says.
David Gosselin and Dylan Skola in the Glass lab, together with Coufal and their collaborators, set out to characterize the molecular characteristics of microglia. They worked with neurosurgeons at UC San Diego to collect brain tissue from 19 patients, all of who were having brain surgery for epilepsy, a brain tumor or a stroke. They isolated microglia from areas of tissue that were unaffected by disease, as well as from mouse brains, and then set out to study the cells. The work was made possible by a multidisciplinary collaboration between bench scientists, bioinformaticians and clinicians.
The team used a variety of molecular and biochemical tests—performed within hours of the cells being collected—to characterize which genes are turned on and off in microglia, how the DNA is marked up by regulatory molecules, and how these patterns change when the cells are cultured.
Microglia, they found, have hundreds of genes that are more highly expressed than other types of macrophages, as well as distinct patterns of gene expression compared to other types of brain cells. After the cells were cultured, however, the gene patterns of the microglia began to change. Within just six hours, more than 2,000 genes had their expression turned down by at least fourfold. The results underscore how dependent microglia are on their surroundings in the brain, and why researchers have struggled to culture them.
Next, the researchers analyzed whether any of the genes that were upregulated in microglia compared to other cells had been previously implicated in disease. Genes linked to a variety of neurodegenerative and psychiatric diseases, they found, were highly expressed in microglia.
“A really high proportion of genes linked to multiple sclerosis, Parkinson’s and schizophrenia are much more highly expressed in microglia than the rest of the brain,” says Coufal. “That suggests there’s some kind of link between microglia and the diseases.”
For Alzheimer’s, more than half of the genes known to affect a person’s risk of developing the disease were expressed more highly in microglia than other brain cells.
In mice, however, many of the disease genes weren’t as highly expressed in microglia. “That tells us that maybe mice aren’t the best model organisms for some of these diseases,” Coufal says.
More work is needed to understand exactly how microglia may be altered in people with diseases, but the new molecular profile of microglia offers a way for researchers to begin trying to better culture the cells, or coax stem cells to develop into microglia for future studies.
Other researchers on the study were Baptiste Jaeger, Carolyn O’Connor, Conor Fitzpatrick, Monique Pena, and Amy Adair of the Salk Institute; Inge Holtman, Johannes Schlachetzki, Eniko Sajti, Martina Pasillas, David Gona, and Michael Levy of the University of California San Diego; and Richard Ransohoff of Biogen.
Funding: The work and the researchers involved were supported by grants from the Larry L. Hillblom Foundation, National Institutes of Health, Canadian Institute of Health Research, Multiple Sclerosis Society of Canada, University of California San Diego, Dutch MS Research Foundation, the Gemmy and Mibeth Tichelaar Foundation, the DFG, the JPB Foundation, Dolby Family Ventures, The Paul G. Allen Family Foundation, the Engman Foundation, the Ben and Wanda Hildyard Chair in Hereditary Diseases.
Source: Salk Institute
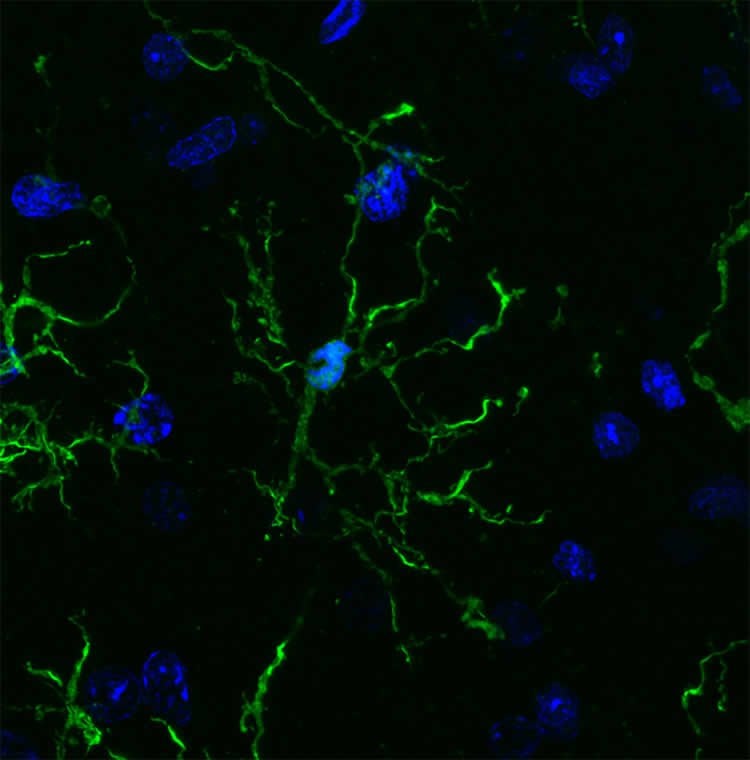
Image Source: NeuroscienceNews.com image is credited to Nicole Coufal.
Original Research: Abstract for “An environment-dependent transcriptional network specifies human microglia identity” by David Gosselin, Dylan Skola, Nicole G. Coufal, Inge R. Holtman, Johannes C. M. Schlachetzki, Eniko Sajti, Baptiste N. Jaeger, Carolyn O’Connor, Conor Fitzpatrick, Martina P. Pasillas, Monique Pena, Amy Adair, David G. Gonda, Michael L. Levy, Richard M. Ransohoff, Fred H. Gage, and Christopher K. Glass in Science. Published online May 25 2017 doi:10.1126/science.aal3222
[cbtabs][cbtab title=”MLA”]Salk Institute “Immune Cells Linked to Schizophrenia, Parkinson’s and Alzheimer’s.” NeuroscienceNews. NeuroscienceNews, 31 May 2017.
<https://neurosciencenews.com/alzheimers-schizophrenia-immune-cells-6810/>.[/cbtab][cbtab title=”APA”]Salk Institute (2017, May 31). Immune Cells Linked to Schizophrenia, Parkinson’s and Alzheimer’s. NeuroscienceNew. Retrieved May 31, 2017 from https://neurosciencenews.com/alzheimers-schizophrenia-immune-cells-6810/[/cbtab][cbtab title=”Chicago”]Salk Institute “Immune Cells Linked to Schizophrenia, Parkinson’s and Alzheimer’s.” https://neurosciencenews.com/alzheimers-schizophrenia-immune-cells-6810/ (accessed May 31, 2017).[/cbtab][/cbtabs]
Abstract
An environment-dependent transcriptional network specifies human microglia identity
Microglia play essential roles in central nervous system (CNS) homeostasis and influence diverse aspects of neuronal function. However, the transcriptional mechanisms that specify human microglia phenotypes are largely unknown. We examined the transcriptomes and epigenetic landscapes of human microglia isolated from surgically resected brain tissue ex vivo and following transition to an in vitro environment. Transfer to a tissue culture environment results in rapid and extensive downregulation of microglia-specific genes that are induced in primitive mouse macrophages following migration into the fetal brain. Substantial subsets of these genes exhibit altered expression in neurodegenerative and behavioral diseases and are associated with non-coding risk variants. These findings reveal an environment-dependent transcriptional network specifying microglia-specific programs of gene expression and facilitate efforts to understand the roles of microglia in human disease.
“An environment-dependent transcriptional network specifies human microglia identity” by David Gosselin, Dylan Skola, Nicole G. Coufal, Inge R. Holtman, Johannes C. M. Schlachetzki, Eniko Sajti, Baptiste N. Jaeger, Carolyn O’Connor, Conor Fitzpatrick, Martina P. Pasillas, Monique Pena, Amy Adair, David G. Gonda, Michael L. Levy, Richard M. Ransohoff, Fred H. Gage, and Christopher K. Glass in Science. Published online May 25 2017 doi:10.1126/science.aal3222






