Research sheds light the neural structure that controls our sleep, eating habits, hormones and more.
What’s that old saying about Mussolini? Say what you will but he made the trains run on time. Well, the suprachiasmatic nucleus — SCN for short — makes everything in the body run on time. The SCN is the control center for our internal genetic clock, the circadian rhythms which regulate everything from sleep to hunger, insulin sensitivity, hormone levels, body temperature, cell cycles and more.
The SCN has been studied extensively but the underlying structure of its neural network has remained a mystery.
Now, researchers from the Harvard John A. Paulson School of Engineering and Applied Sciences (SEAS), the University of California Santa Barbara, and Washington University in St. Louis have shown for the first time how neurons in the SCN are connected to each other, shedding light on this vital area of the brain. Understanding this structure — and how it responds to disruption — is important for tackling illnesses like diabetes and posttraumatic stress disorder. The scientists have also found that disruption to these rhythms such as shifts in work schedules or blue light exposure at night can negatively impact overall health.
The research was recently published in the Proceedings of the National Academies of Science (PNAS).
“The SCN has been so challenging to understand because the cells within it are incredibly noisy,” said John Abel, first author of the paper and graduate student at SEAS. “There are more than 20,000 neurons in the SCN, each of which not only generates their own autonomous circadian oscillations but also communicates with other neurons to maintain stable phase lengths and relationships. We were able to cut through that noise and figure out which cells share information with each other.”
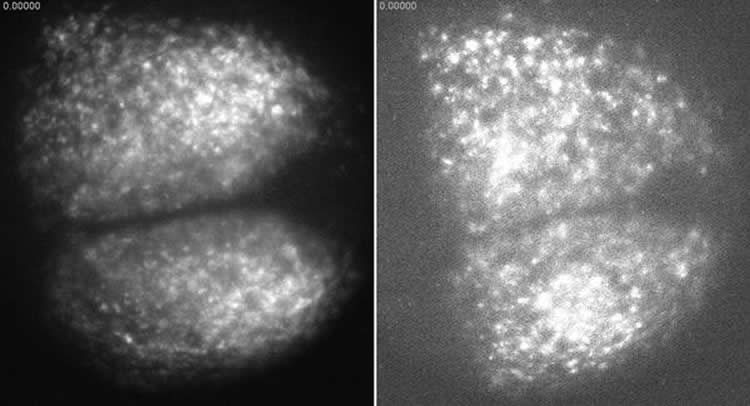
The SCN looks like a miniature brain, with two hemispheres, inside the hypothalamus. It receives light cues from the retina to help it keep track of time and reset when necessary. When functioning probably, the neurons inside both hemispheres oscillate in a synchronized pattern.
In order to understand the structure of the network, Abel and the team had to disrupt that pattern. The researchers used a potent neurotoxin commonly found in pufferfish to desynchronize the neurons in each hemisphere, turning the steady, rhythmic pulse of oscillations into a cacophony of disconnected beats. The team then removed the toxin and observed the network as it reestablished communication, using information theory to figure out which cells had to communicate to resynchronize the whole network.
“It’s like trying to figure out if a group of people are friends without being able to look at their phone calls or their text messages,” Abel said. “In a large group of other people, you might not be able to tell who is in contact with each other, but if a certain group shows up together at a party, you can probably assume they’re friends because they show similar behavior.”
By observing the SCN at single-cell resolution, Abel and the team identified a core group of very friendly neurons in the center of each hemisphere that share a lot of information during resynchronization. They also observed dense connections between the hubs of each hemisphere. The neurons outside these central hubs, in the area called the shell, behaved more like acquaintances than friends, sharing little information amongst themselves.
“We were surprised to find that the shell lacked a functionally connected cluster of neurons,” said Abel. “We’ve known that exposure to an artificially long day can split the SCN into core and shell phase clusters which oscillate out of sync with each other. We’ve assumed that the neurons in the shell communicated to synchronize that rhythm but our research suggests that phase clustering in the shell is actually mediated by the core neurons.”
Previous research also assumed that the core SCN was dominant only due to its role in receiving light cues from the eyes. By using the neurotoxin to disrupt circadian rhythms, Abel and the team demonstrated that the core is the key to resynchronization even without light cues.
“For the last 15 years our group has been studying the complex control mechanisms that are responsible for the generation of robust circadian rhythms in the brain,” said Frank Doyle, the John A. Paulson Dean and John A. & Elizabeth S. Armstrong Professor of Engineering & Applied Sciences, who co-authored the paper. “This work brings us one step closer to reverse engineering those paradigms by elucidating the topology of communication amongst neurons, thus demonstrating the importance of a systems perspective to link genes to cells to the SCN tissue.”
The research was coauthored by Kirsten Meeker, Peter St. John, Benjamin Bales and Linda Petzold of UC Santa Barbara and Thomas Wang, Daniel Granados-Fuentes and Eric Herzog, of Washington University.
Funding: It was funded by the National Institute of Health and the US Army Research Office.
Source: Leah Burrows – Harvard
Image Source: The image is credited to Doyle Lab.
Original Research: Abstract for “Functional network inference of the suprachiasmatic nucleus” by John H. Abel, Kirsten Meeker, Daniel Granados-Fuentes, Peter C. St. John, Thomas J. Wang, Benjamin B. Bales, Francis J. Doyle III, Erik D. Herzog, and Linda R. Petzold in PNAS. Published online April 4 2016 doi:10.1073/pnas.1521178113
Abstract
Functional network inference of the suprachiasmatic nucleus
In the mammalian suprachiasmatic nucleus (SCN), noisy cellular oscillators communicate within a neuronal network to generate precise system-wide circadian rhythms. Although the intracellular genetic oscillator and intercellular biochemical coupling mechanisms have been examined previously, the network topology driving synchronization of the SCN has not been elucidated. This network has been particularly challenging to probe, due to its oscillatory components and slow coupling timescale. In this work, we investigated the SCN network at a single-cell resolution through a chemically induced desynchronization. We then inferred functional connections in the SCN by applying the maximal information coefficient statistic to bioluminescence reporter data from individual neurons while they resynchronized their circadian cycling. Our results demonstrate that the functional network of circadian cells associated with resynchronization has small-world characteristics, with a node degree distribution that is exponential. We show that hubs of this small-world network are preferentially located in the central SCN, with sparsely connected shells surrounding these cores. Finally, we used two computational models of circadian neurons to validate our predictions of network structure. pathways, could lead to repurposed therapeutics.
“Functional network inference of the suprachiasmatic nucleus” by John H. Abel, Kirsten Meeker, Daniel Granados-Fuentes, Peter C. St. John, Thomas J. Wang, Benjamin B. Bales, Francis J. Doyle III, Erik D. Herzog, and Linda R. Petzold in PNAS. Published online April 4 2016 doi:10.1073/pnas.1521178113