Summary: Researchers have identified a gene network that regulates the development of motor neurons during embryonic development.
Source: UCLA.
UCLA researchers have discovered the inner workings of a gene network that regulates the development of spinal motor neurons in the growing chicken and mouse embryo. The research also answers a long-standing question about why motor neurons, the nerve cells of the spinal cord that control muscle movement, form much faster than other types of neurons.
The study was published in the journal PLOS Biology by co-senior author Bennett Novitch, member of the Eli and Edythe Broad Center of Regenerative Medicine and Stem Cell Research at UCLA and collaborators from the Francis Crick Institute in London, U.K. It has important implications for facilitating the production of motor neurons from stem cells in the lab. Stem cell-derived motor neurons could be used to repair the diseased or damaged spinal cord and to study neurodegenerative diseases such as amyotrophic lateral sclerosis (also known as Lou Gehrig’s disease) and spinal muscular atrophy.
“This study provides an unprecedented detailed view of how embryos produce the different cell types found in the mature spinal cord,” said Novitch, UCLA’s Ethel Scheibel Professor of Neurobiology. “The new experimental techniques we used, combined with collaborative efforts of biologists and computer scientists, are allowing us to gain new insight into the exquisite accuracy of embryonic development.”
During embryonic development of the spinal cord, different types of nerve cells are formed from precursor cells, called neural progenitors. It has been known for a long time that different types of neurons form at differing speeds during development, with motor neurons forming faster than the other types of nerve cells that populate the spinal cord. However, researchers didn’t know why or how motor neurons develop this way.
The research team used the latest molecular techniques to assess how the activity of genes change as neurons form. This was achieved using an approach called single cell transcriptional profiling, which allows the activity of all genes in individual cells to be measured simultaneously. This gave the team snapshots of global gene activity in about 200 cells that were in the process of becoming motor neurons. To analyze these data, the team developed custom computer software to reconstruct how gene activity changes as motor neurons form.
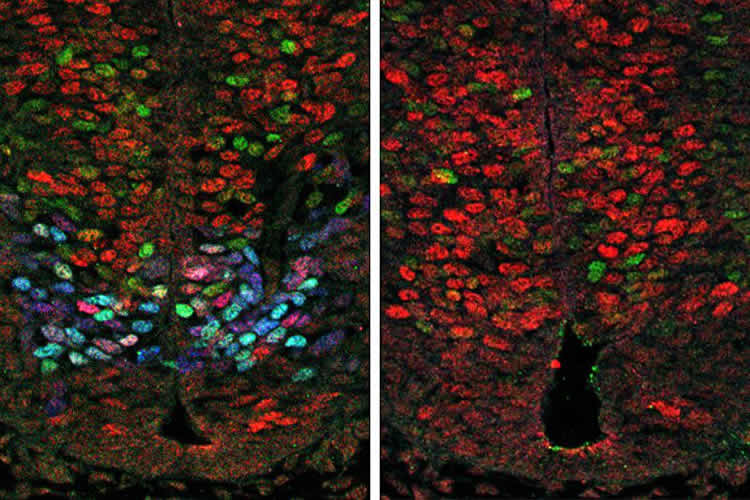
The analysis suggested that a protein produced by a gene called Olig2, which is only expressed in the neural progenitors that create motor neurons, promotes motor neuron formation by interfering with the activity of several members of another gene family—the Hes genes. The Hes genes are known to prevent the development of neurons. The researchers confirmed Olig2’s role in promoting motor neuron formation by increasing or blocking the function of Olig2 in the spinal cords of developing mouse and chicken embryos, as well as during motor neuron formation in mouse embryonic stem cells.
“During embryonic development, the nervous system is constructed following a precise blueprint, with key parts forming at precise times and places, and appropriate numbers. Our research sheds light into how this process is orchestrated specifically for motor neuron development,” said Novitch.
Funding: The research was funded by the National Institute for Neurological Disease and Stroke, the March of Dimes Foundation, the Whitehall Foundation and the UCLA Broad Stem Cell Research Center.
Source: Mirabai Vogt-James – UCLA
Publisher: Organized by NeuroscienceNews.com.
Image Source: NeuroscienceNews.com image is credited to the researchers.
Original Research: Open access research in PLOS Biology.
doi:10.1371/journal.pbio.2003127
[cbtabs][cbtab title=”MLA”]UCLA “Gene Network That Regulates Motor Neuron Formation During Embryonic Development Identified.” NeuroscienceNews. NeuroscienceNews, 5 February 2018.
<https://neurosciencenews.com/motor-neuron-development-8431/>.[/cbtab][cbtab title=”APA”]UCLA (2018, February 5). Gene Network That Regulates Motor Neuron Formation During Embryonic Development Identified. NeuroscienceNews. Retrieved February 5, 2018 from https://neurosciencenews.com/motor-neuron-development-8431/[/cbtab][cbtab title=”Chicago”]UCLA “Gene Network That Regulates Motor Neuron Formation During Embryonic Development Identified.” https://neurosciencenews.com/motor-neuron-development-8431/ (accessed February 5, 2018).[/cbtab][/cbtabs]
Abstract
Olig2 and Hes regulatory dynamics during motor neuron differentiation revealed by single cell transcriptomics
During tissue development, multipotent progenitors differentiate into specific cell types in characteristic spatial and temporal patterns. We addressed the mechanism linking progenitor identity and differentiation rate in the neural tube, where motor neuron (MN) progenitors differentiate more rapidly than other progenitors. Using single cell transcriptomics, we defined the transcriptional changes associated with the transition of neural progenitors into MNs. Reconstruction of gene expression dynamics from these data indicate a pivotal role for the MN determinant Olig2 just prior to MN differentiation. Olig2 represses expression of the Notch signaling pathway effectors Hes1 and Hes5. Olig2 repression of Hes5 appears to be direct, via a conserved regulatory element within the Hes5 locus that restricts expression from MN progenitors. These findings reveal a tight coupling between the regulatory networks that control patterning and neuronal differentiation and demonstrate how Olig2 acts as the developmental pacemaker coordinating the spatial and temporal pattern of MN generation.