Summary: A study in Nature Neuroscience reveals mutations that occur after conception could play an important role in autism. The study reports post-zygotic mutations occur disproportionately in genes expressed in the amygdala in those on the autism spectrum.
Source: Boston Children’s Hospital.
Large study maps mutations arising after conception; many affect the amygdala.
A study of nearly 6,000 families, combining three genetic sequencing technologies, finds that mutations that occur after conception play an important role in autism. A team led by investigators at Boston Children’s Hospital and the Broad Institute of MIT and Harvard reports the findings today in Nature Neuroscience.
Over the past decade, mutations to more than 60 different genes have been linked with autism spectrum disorder (ASD), including spontaneous, non-inherited (de novo) mutations. But much of autism still remains unexplained. The study, led by first author Elaine Lim, PhD, of Boston Children’s, and senior author Christopher Walsh, MD, PhD, of Boston Children’s and the Broad Institute, delved into an emerging category of de novo mutations: those found in only a subset of our cells.
De novo mutations can occur in a parent’s sperm or egg, or they can occur after fertilization, arising in an embryonic cell. These are known as post-zygotic mutations or PZMs (also known as somatic mutations). The later PZMs occur during embryonic development, the fewer cells will carry them, making them harder to detect.
“If the mutation is in a very small fraction of all cells, it will be missed by whole-exome sequencing,” says Lim.
Finding post-zygotic mutations
To find PZMs, Lim, Walsh and colleagues obtained whole-exome sequencing data previously gathered from 5,947 families, courtesy of the Simons Foundation Autism Research Initiative (SFARI) Simplex Collection, the Autism Sequencing Consortium and Autism Speaks. They then resequenced some of the DNA from these children using three independent sequencing technologies in parallel.
Based on their findings, they classified 7.5 percent of ASD subjects’ de novo mutations as PZMs. Of these, 83 percent had not been picked up in the original analysis of their genome sequence.
Some PZMs affected genes already known to be linked to autism or other neurodevelopmental disorders (such as SCN2A, HNRNPU and SMARCA4), but sometimes affected these genes in different ways. Many others were in genes known to be active in brain development (such as KLF16 and MSANTD2) but not previously associated with ASD.
The connection of these genes to autism may have been missed before because the earlier studies focused on mutations that knocked down gene function. “Some of the postzygotic mutations we found represented a gain of function, not a loss of function,” says Lim, who is also affiliated with the Wyss Institute for Biologically Inspired Engineering.
The amygdala and autism
To estimate the developmental timing of the mutations and the brain regions affected, Lim, Walsh and colleagues compared their sequencing data, which mostly came from blood DNA samples, with publicly available gene expression data from brain autopsies representing different ages (prenatal through adult) via the BrainSpan Project.
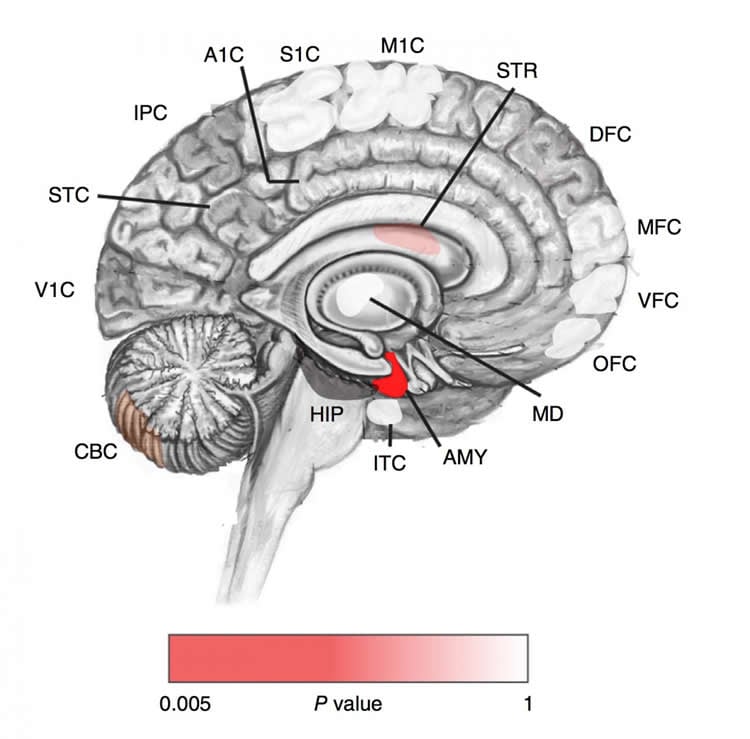
“By overlapping the data, we can start to map where in the brain these genes are expressed and when the mutations occurred during development,” says Lim.
These analyses showed that PZMs in the subjects with ASD occur disproportionately in genes expressed in the amygdala. “This was exciting to us, in that the amygdala has been proposed as an important region of the brain in autism,” says Lim.
Our dynamically developing brains
Overall, the work adds to evidence that complex brain disorders, such as epilepsy, intellectual disability, schizophrenia and brain malformations, can arise from non-inherited mutations that occur at some point during prenatal development.
“We have known that PZMs are an important cause of epilepsy, but this work provides the best evidence so far that they are relevant to autism as well,” says Walsh, who is also an Investigator of the Howard Hughes Medical Institute. “So it is now exciting to consider what other psychiatric conditions might have a role for PZMs.”
Funding: Supporters of the study include the Simons Foundation, the National Institutes of Health (R01MH083565, RC2MH089952, U01MH106883 and others) and the Howard Hughes Medical Institute.
Source: Keri Stedman – Boston Children’s Hospital
Image Source: NeuroscienceNews.com image is credited to Mohammed Uddin.
Original Research: Abstract for “Rates, distribution and implications of postzygotic mosaic mutations in autism spectrum disorder” by Elaine T Lim, Mohammed Uddin, Silvia De Rubeis, Yingleong Chan, Anne S Kamumbu, Xiaochang Zhang, Alissa M D’Gama, Sonia N Kim, Robert Sean Hill, Arthur P Goldberg, Christopher Poultney, Nancy J Minshew, Itaru Kushima, Branko Aleksic, Norio Ozaki, Mara Parellada, Celso Arango, Maria J Penzol, Angel Carracedo, Alexander Kolevzon, Christina M Hultman, Lauren A Weiss, Menachem Fromer, Andreas G Chiocchetti, Christine M Freitag, Autism Sequencing Consortium, George M Church, Stephen W Scherer, Joseph D Buxbaum & Christopher A Walsh in Nature Neuroscience. Published online July 17 2017 doi:10.1038/nn.4598
[cbtabs][cbtab title=”MLA”]Boston Children’s Hospital “Late Breaking Mutations May Play an Important Role in Autism.” NeuroscienceNews. NeuroscienceNews, 17 July 2017.
<https://neurosciencenews.com/late-mutation-autism-7092/>.[/cbtab][cbtab title=”APA”]Boston Children’s Hospital (2017, July 17). Late Breaking Mutations May Play an Important Role in Autism. NeuroscienceNew. Retrieved July 17, 2017 from https://neurosciencenews.com/late-mutation-autism-7092/[/cbtab][cbtab title=”Chicago”]Boston Children’s Hospital “Late Breaking Mutations May Play an Important Role in Autism.” https://neurosciencenews.com/late-mutation-autism-7092/ (accessed July 17, 2017).[/cbtab][/cbtabs]
Abstract
Rates, distribution and implications of postzygotic mosaic mutations in autism spectrum disorder
We systematically analyzed postzygotic mutations (PZMs) in whole-exome sequences from the largest collection of trios (5,947) with autism spectrum disorder (ASD) available, including 282 unpublished trios, and performed resequencing using multiple independent technologies. We identified 7.5% of de novo mutations as PZMs, 83.3% of which were not described in previous studies. Damaging, nonsynonymous PZMs within critical exons of prenatally expressed genes were more common in ASD probands than controls (P < 1 × 10−6), and genes carrying these PZMs were enriched for expression in the amygdala (P = 5.4 × 10−3). Two genes (KLF16 and MSANTD2) were significantly enriched for PZMs genome-wide, and other PZMs involved genes (SCN2A, HNRNPU and SMARCA4) whose mutation is known to cause ASD or other neurodevelopmental disorders. PZMs constitute a significant proportion of de novo mutations and contribute importantly to ASD risk.
“Rates, distribution and implications of postzygotic mosaic mutations in autism spectrum disorder” by Elaine T Lim, Mohammed Uddin, Silvia De Rubeis, Yingleong Chan, Anne S Kamumbu, Xiaochang Zhang, Alissa M D’Gama, Sonia N Kim, Robert Sean Hill, Arthur P Goldberg, Christopher Poultney, Nancy J Minshew, Itaru Kushima, Branko Aleksic, Norio Ozaki, Mara Parellada, Celso Arango, Maria J Penzol, Angel Carracedo, Alexander Kolevzon, Christina M Hultman, Lauren A Weiss, Menachem Fromer, Andreas G Chiocchetti, Christine M Freitag, Autism Sequencing Consortium, George M Church, Stephen W Scherer, Joseph D Buxbaum & Christopher A Walsh in Nature Neuroscience. Published online July 17 2017 doi:10.1038/nn.4598






