Summary: Genetics and microenvironment influence the frequency of glioblastoma cells. Researchers provide a new blueprint for glioblastoma, integrating the malignant cell programs, cancer cell plasticity, and modulation by genetic drivers. The findings shed light on why this form of cancer is so hard to treat effectively.
Source: Broad Institute
Glioblastoma is an incurable brain cancer that kills most patients less than two years after diagnosis. The disease is difficult to treat largely because tumors each contain multiple kinds of cells. The aggressive brain cancer can also vastly differ between patients, so much so that researchers debate whether glioblastoma should be considered a single disease. A new study may help clarify what drives this important heterogeneity and makes glioblastoma so deadly.
The researchers profiled gene expression in more than 24,000 tumor cells from 20 adult and eight pediatric glioblastoma patients and also analyzed glioblastoma models in the lab. They found four glioblastoma cellular states, which each has a unique gene expression program and together help account for the large variation in the disease. The scientists then used the single-cell data to reanalyze glioblastoma data from The Cancer Genome Atlas and identified genetic alterations associated with each of the four states.
The results, published in Cell, also show that the cells are remarkably flexible or plastic — that is, they can switch between the four states. This shape-shifting ability could help explain why these cancer cells are so difficult to kill with drugs and help inform the development of better therapies for glioblastoma.
The research was led by co-first authors Cyril Neftel, Department of Pathology at Massachusetts General Hospital (MGH) and Broad Institute; Julie Laffy, Weizmann Institute of Science; Mariella Filbin, Department of Pathology at MGH, Broad Institute, and Dana-Farber Institute; Toshiro Hara, Salk Institute for Biological Studies; and co-senior authors Itay Tirosh, Weizmann Institute of Science, and Mario Suvà, Department of Pathology at MGH and Broad Institute.
THE STATE OF THE CELL
Previous studies suggested that glioblastoma samples fall into at least three subtypes: proneural, classical, and mesenchymal, which differ mostly by the genes they express. Researchers have also shown that glioblastoma tumors are almost always made up of more than one subtype, and that the proportions of different subtypes in a tumor can change over time and during treatment. However, most of these studies looked at bulk samples, where genetic material from many tumor cells is combined and sequenced. This method provides a rough idea of the subtypes in a tumor and their relative abundance, but not the finer details.
To better understand glioblastoma states at the cellular level, the researchers examined cells using single-cell RNA sequencing, a suite of genomic techniques that reveal individual cells’ gene expression profiles.
“We are placing the individual cellular states and their associated genetic alterations in their developmental context and understanding what cell types are driving the disease,” said Suvà, who is an institute member in the Broad’s Epigenomics Program, a molecular pathologist in MGH’s Department of Pathology, and a member of the Center for Cancer Research at MGH. He added that this effort is the largest single-cell sequencing study for glioblastoma to date.
The researchers discovered four distinct gene expression programs among the cells, representing four dominant cellular states: neural-progenitor-like, oligodendrocyte-progenitor-like, astrocyte-like, and mesenchymal-like. All four states resemble normal brain cell types to some extent, but with important differences that make the cells cancerous. These states also correlate well with the previously-described subtypes in glioblastoma.
The four states were represented in both adult and pediatric tumors, and the majority of tumors contained cells in every state while the remaining minority had cells representing at least two states. However, the proportions of cell states varied among tumors.
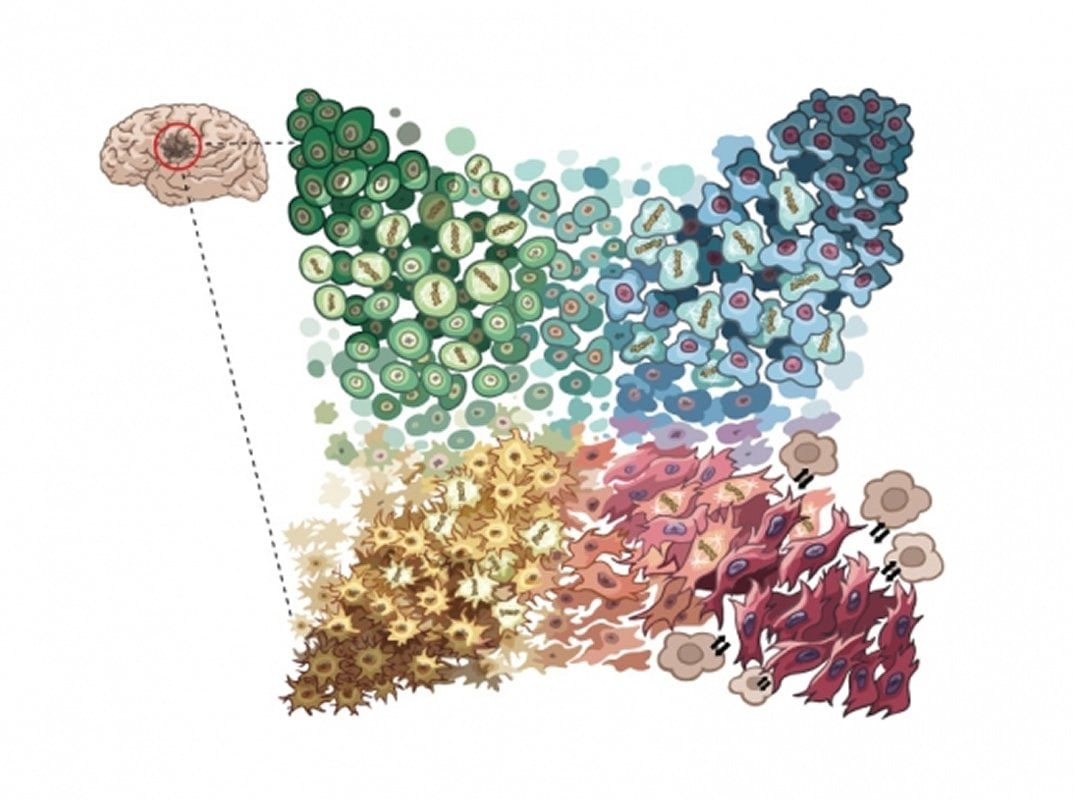
To see whether these differences were due to genetics, the tumor’s surrounding microenvironment, or both, the researchers looked for genes that were associated with tumors containing a high percentage of cells in a particular cell state. They determined that common genetic drivers of glioblastoma (genes including EGFR, PDGFRA, and CDK4) play a role in which state a glioblastoma cell takes and how many cells of that state are in a patient sample. For one of the states, mesenchymal-like, they also found that the microenvironment played an important role.
CROSSING STATE LINES
While most cells seemed to fit clearly into one of the four states, about 15% expressed two programs simultaneously.
“Finding a lot of cells in between two states is strong evidence that those states can transition from one to the other,” Suvà said.
To test this observation, the researchers injected mice with cells derived from a human glioblastoma tumor, with each mouse receiving cells that represented only one state. They found that the resulting tumors contained not just that cell state, but all the other states that were present in the original tumor, and in about the same proportions. This suggests the glioblastoma cells can transition from one state to any of the others. The team came to the same conclusion after using genetic barcoding to follow how individual cells developed and changed in both a mouse glioblastoma model and in mice harboring human glioblastoma cells.
The findings provide clues as to why existing cancer treatments fail to stop glioblastoma growth. “If you understand that this is a disease with multiple states driving it, each one with a corresponding favorite cancer gene, you understand better why targeting one gene at a time has failed so far, and you can dissect the mechanisms of adaptation and resistance to therapies,” said Suvà.
The researchers now plan to study how current therapies affect each of the four states and search for novel ways to target each state.
Other key Broad Institute members involved in this research include Marni Shore, Gilbert Rahme, Bradley Bernstein, and core institute member Aviv Regev. Shore, Rahme, and Bernstein are also members of the Department of Pathology and Center for Cancer Research at MGH and Regev is also a Howard Hughes Medical Institute investigator and a professor of biology at MIT.
Funding: Funding for this study was provided in part by the Sontag Foundation, the Howard Goodman Fellowship at MGH, the Merkin Institute Fellowship at the Broad Institute of MIT and Harvard, the Wang Family Fund, the Smith Family Foundation, the Chan-Zuckerberg Initiative, the Swiss National Science Foundation Sinergia Grant, Alex’s Lemonade Stand, the V Foundation for Cancer Research, the Zuckerman STEM Leadership Program, the Human Frontiers Science Program, and start-up funds from the MGH Department of Pathology and the Weizmann Institute of Science.
Source:
Broad Institute
Media Contacts:
Abbey Bigler – Broad Institute
Image Source:
The image is credited to Sébastien Perroud.
Original Research: Closed access
“An integrative model of cellular states, plasticity and genetics for glioblastoma”. Neftel C, Laffy J, Filbin MG, Hara T, et al.
Cell. doi:10.1016/j.cell.2019.06.024
Abstract
An integrative model of cellular states, plasticity and genetics for glioblastoma
Highlights
• Four cellular states drive glioblastoma malignant cells heterogeneity
• In vivo single-cell lineage tracing supports plasticity between these four states
• Genetics and the microenvironment influence the frequency of cells in each state
• TCGA subtypes reflect the highest-frequency malignant states and the microenvironment
Summary
Diverse genetic, epigenetic, and developmental programs drive glioblastoma, an incurable and poorly understood tumor, but their precise characterization remains challenging. Here, we use an integrative approach spanning single-cell RNA-sequencing of 28 tumors, bulk genetic and expression analysis of 401 specimens from the The Cancer Genome Atlas (TCGA), functional approaches, and single-cell lineage tracing to derive a unified model of cellular states and genetic diversity in glioblastoma. We find that malignant cells in glioblastoma exist in four main cellular states that recapitulate distinct neural cell types, are influenced by the tumor microenvironment, and exhibit plasticity. The relative frequency of cells in each state varies between glioblastoma samples and is influenced by copy number amplifications of the CDK4, EGFR, and PDGFRA loci and by mutations in the NF1 locus, which each favor a defined state. Our work provides a blueprint for glioblastoma, integrating the malignant cell programs, their plasticity, and their modulation by genetic drivers.






