Summary: Researchers report FMR1 and ZC3H14 proteins both regulate mRNA, which may explain their importance in memory and learning.
Source: Emory Health Sciences.
RNA-binding proteins involved in inherited intellectual disability.
We can learn a lot about somebody from the friends they hang out with. This applies to people and also to genes and proteins. Emory scientists have been investigating a gene that we will call – spoiler alert –“Friend of fragile X.”
Fragile X syndrome is the most common inherited form of intellectual disability, studied by research teams around the world with drug discovery and clinical trials in mind. It is caused by a disruption of the gene FMR1.
In an independent form of inherited intellectual disability found in a small number of Iranian families, a gene called ZC3H14 is mutated. Two papers from Ken Moberg, PhD, associate professor of cell biology, Anita Corbett, PhD, professor of biology and colleagues show that FMR1 and ZC3H14 are, in effect, friends.
The findings provide new insight into the function of FMR1 as well as ZC3H14; the evidence comes from experiments performed in fruit flies and mice. The most recent paper is in the journal Cell Reports, published August 8, 2017.
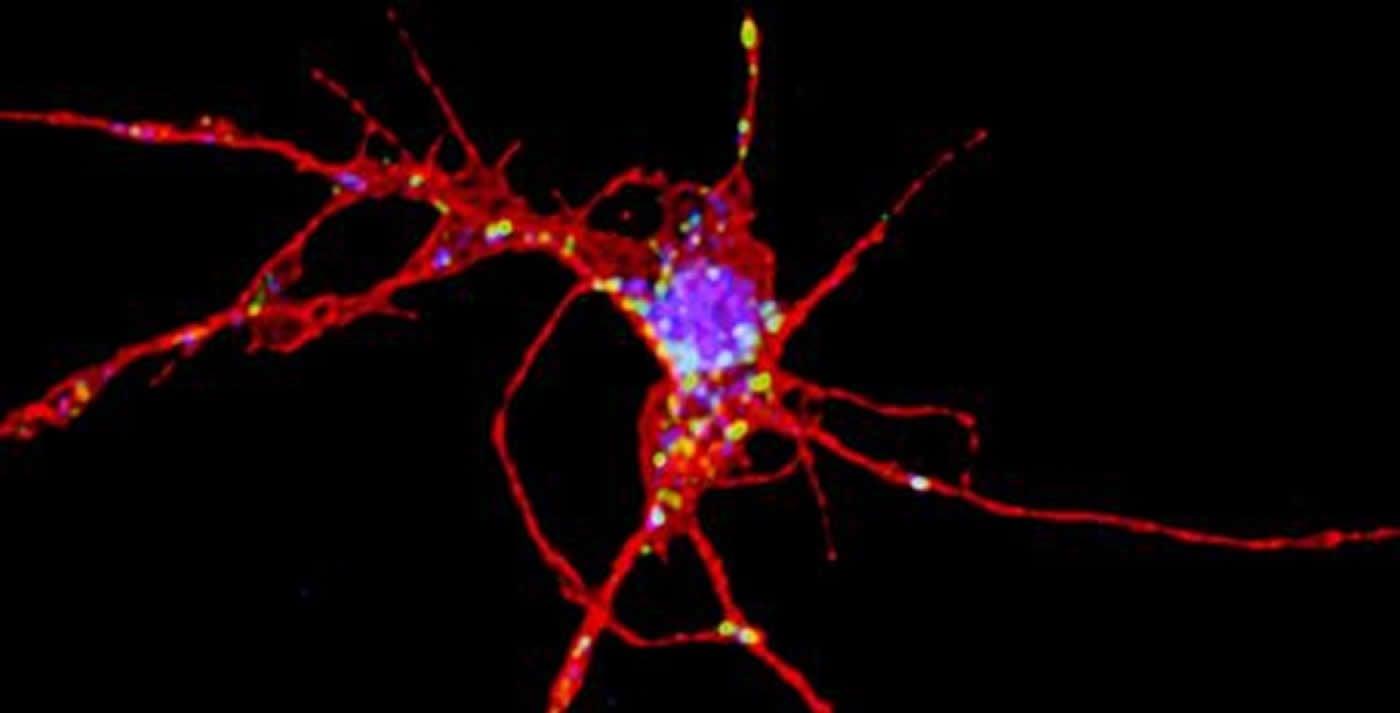
The scientists found that the proteins encoded by FMR1 and ZC3H14 stick together in cells and they hang out in the same places. The two proteins have related functions: they both regulate messenger RNA in neurons, which explains their importance for learning and memory.
The fragile X protein (FMRP) was known to control protein production in response to signals arriving in neurons, but the Cell Reports paper shows that FMRP is also regulating the length of “tails” attached to messenger RNAs – something scientists did not realize, even after years of studying FMRP and fragile X, Moberg says.
In fruit flies, ZC3H14 is called dNab2. Mutations in this gene were previously shown to hinder flies’ flying and climbing, as well as impair memory formation during courtship.
In the new paper, the researchers show that dNab2 mutant flies also have problems learning to avoid situations when an unpleasant smell is present. In this learning task, dNab2 and the fly version of FMR1 are working together, the researchers showed. However, the sets of RNAs they regulate do not overlap completely.
Several labs at Emory contributed to the research, including those of human genetics chair Stephen Warren, PhD, cell biology chair Gary Bassell, PhD and James Zheng, PhD. The first author of the Cell Reports paper is Rick Bienkowski, PhD, a former Genetics and Molecular Biology graduate student.
“We could not have done this work without the strength of the neuroscience and especially, fragile X, community at Emory,” Moberg says.
Corbett and Moberg’s collaboration on dNab2/ZC3H14 was accelerated in 2009 when German and Iranian researchers contacted them about the link to human intellectual disability. It also pushed Corbett, who had first examined Nab2’s function in yeast, into the realm of knockout mice.
Her lab’s research on mice lacking ZC3H14, conducted with neuroscientists Paul Garcia, MD, PhD, and Andrew Escayg, PhD, was published in June in the journal Human Molecular Genetics.
Similar to the mutant flies, the mice display impaired working memory and altered brain development. Studies that combine the mutant mice and flies will reveal more about the functions of both RNA-binding proteins in learning and memory, Corbett says.
Funding: Research in Moberg and Corbett’s labs was supported by the National Institute of Mental Health (MH10730501).
Source: Quinn Eastman – Emory Health Sciences
Image Source: NeuroscienceNews.com image is credited to Ken Moberg.
Original Research: Full open access research for “The Conserved, Disease-Associated RNA Binding Protein dNab2 Interacts with the Fragile X Protein Ortholog in Drosophila Neurons” by Rick S. Bienkowski, Ayan Banerjee, J. Christopher Rounds, Jennifer Rha, Omotola F. Omotade, Christina Gross, Kevin J. Morris, Sara W. Leung, ChangHui Pak, Stephanie K. Jones, Michael R. Santoro, Stephen T. Warren, James Q. Zheng, Gary J. Bassell, Anita H. Corbett, and Kenneth H. Moberg in Cell Reports. Published online August 8 2017 doi:10.1016/j.celrep.2017.07.038
[cbtabs][cbtab title=”MLA”]Emory Health Sciences “Insight into Learning via ‘Friend of Fragile X’ Gene.” NeuroscienceNews. NeuroscienceNews, 9 August 2017.
<https://neurosciencenews.com/fragile-x-learning-7269/>.[/cbtab][cbtab title=”APA”]Emory Health Sciences (2017, August 9). Insight into Learning via ‘Friend of Fragile X’ Gene. NeuroscienceNew. Retrieved August 9, 2017 from https://neurosciencenews.com/fragile-x-learning-7269/[/cbtab][cbtab title=”Chicago”]Emory Health Sciences “Insight into Learning via ‘Friend of Fragile X’ Gene.” https://neurosciencenews.com/fragile-x-learning-7269/ (accessed August 9, 2017).[/cbtab][/cbtabs]
Abstract
The Conserved, Disease-Associated RNA Binding Protein dNab2 Interacts with the Fragile X Protein Ortholog in Drosophila Neurons
Highlights
•dNab2 is the fly ortholog of a human RBP lost in inherited intellectual disability
•A cytoplasmic pool of dNab2 interacts with the fragile X homolog dFMRP
•dNab2 regulates the CamKII mRNA and supports memory with dFMRP
•dFMRP and dNab2 both restrict poly(A) length of neuronal mRNAs
Summary
The Drosophila dNab2 protein is an ortholog of human ZC3H14, a poly(A) RNA binding protein required for intellectual function. dNab2 supports memory and axon projection, but its molecular role in neurons is undefined. Here, we present a network of interactions that links dNab2 to cytoplasmic control of neuronal mRNAs in conjunction with the fragile X protein ortholog dFMRP. dNab2 and dfmr1 interact genetically in control of neurodevelopment and olfactory memory, and their encoded proteins co-localize in puncta within neuronal processes. dNab2 regulates CaMKII, but not futsch, implying a selective role in control of dFMRP-bound transcripts. Reciprocally, dFMRP and vertebrate FMRP restrict mRNA poly(A) tail length, similar to dNab2/ZC3H14. Parallel studies of murine hippocampal neurons indicate that ZC3H14 is also a cytoplasmic regulator of neuronal mRNAs. Altogether, these findings suggest that dNab2 represses expression of a subset of dFMRP-target mRNAs, which could underlie brain-specific defects in patients lacking ZC3H14.
“The Conserved, Disease-Associated RNA Binding Protein dNab2 Interacts with the Fragile X Protein Ortholog in Drosophila Neurons” by Rick S. Bienkowski, Ayan Banerjee, J. Christopher Rounds, Jennifer Rha, Omotola F. Omotade, Christina Gross, Kevin J. Morris, Sara W. Leung, ChangHui Pak, Stephanie K. Jones, Michael R. Santoro, Stephen T. Warren, James Q. Zheng, Gary J. Bassell, Anita H. Corbett, and Kenneth H. Moberg in Cell Reports. Published online August 8 2017 doi:10.1016/j.celrep.2017.07.038