Histone demethylase = epigenetic eraser.
Right after fertilization, embryos at the earliest stages of development tell their genes: “Forget what it was like in the sperm or egg where you came from.”
When the process of epigenetic reprogramming is defective in mouse development, the consequences in adulthood can include abnormal repetitive behaviors, scientists have shown.
Their findings are published online in the journal eLife.
“Our results demonstrate how defects in reprogramming may influence the development of altered behaviors, or even complex psychiatric disorders,” says co-senior author David Katz, PhD, assistant professor of cell biology at Emory University School of Medicine.
After fertilization, the enzyme KDM1A (lysine specific demethylase 1A) appears to act as an epigenetic eraser, wiping away information carried on histones, the spool-like proteins that package DNA. KDM1A removes histone methylation, a chemical modification that shapes the activity of nearby genes.
Katz and graduate student Jadiel Wasson created genetically engineered mice that have KDM1A missing from their mouse oocytes (or egg cells), but present later in development. They teamed up with Todd MacFarlan, PhD, previously at the Salk Institute and now at the National Institute of Child Health and Human Development, to examine several mouse strains with alterations in the KDM1A gene.
While these mice were created by genetic engineering techniques, Katz says they may simulate other disruptions of the reprogramming process after fertilization. Those disruptions might come from genetic changes or environmental influences on oocytes such as hormones or parental age, he says.
“These mice have functioning KDM1A later on in development, because they inherit a good copy of the gene from their fathers,” Macfarlan says. “But it’s not there at a critical stage — what we call the maternal-to-zygote transition.”
In the situation when egg cells are missing the enzyme completely, much of the pre-fertilization information present on histones persists after fertilization. That means thousands of egg- and sperm-specific genes are turned on that shouldn’t be, and a similar number needed for embryonic development are turned off.
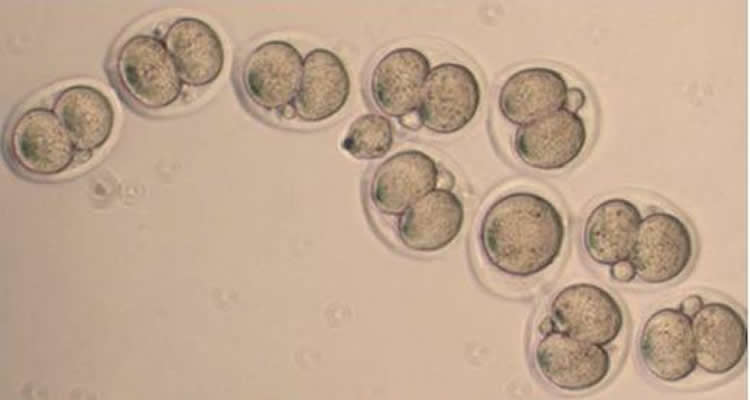
“Usually, that’s lethal,” Katz says. “The fertilized egg can’t tolerate it at all, and is unable to divide beyond the two cell stage.”
In contrast, in some of the mice Katz and Wasson studied, a bit of KDM1A enzyme remained in the oocytes. Although most of the resulting mice died in utero or right after birth, a few survived to adulthood. They looked like normal mice, but they behaved strangely.
Wasson says one aspect of their behavior became apparent when more than one was in a cage at the same time: they ground up much of their food and incorporated it into their bedding. The KDM1A-reduced mice also displayed excessive scratching and digging. When tested on marble burying, a task which has been used to gauge obsessive-compulsive behavior in other genetically modified mice, these mice were even more avid marble buriers than had been observed elsewhere.
What accounts for the strange behavior? Wasson and Katz are beginning to examine changes in the brains of the KDM1A-reduced mice. One possible contributor is changes they observed in imprinted genes. With imprinted genes, one copy is silenced depending on whether it comes from the mother or father; these patterns are disrupted in the KDM1A-reduced mice.
“The most likely scenario, and one that we are pursuing, is that the disruptions in gene expression ultimately influence the wiring of the brain,” Katz says. “This could be via imprinting, but is just as likely to be via disruption of normal neuronal development.”
To study KDM1A’s role in brain development, Katz and Macfarlan are planning to create and examine mice that have a less drastic reduction of the KDM1A enzyme in oocytes, so that more survive to adulthood.
Funding: The research was supported by the National Science Foundation (IOS1354998) and Eunice Kennedy Shriver National Institute of Child Health and Human Development DIR grant HD008933. Wasson is part of the Biochemistry, Cell and Developmental Biology graduate program at Emory.
Source: Quinn Eastman – Emory University
Image Source: The image is credited to Jadiel Wasson
Original Research: Abstract for “Maternally provided LSD1/KDM1A enables the maternal-to-zygotic transition and prevents defects that manifest postnatally” by Jadiel A Wasson, Ashley K Simon, Dexter A Myrick, Gernot Wolf, Shawn Driscoll, Samuel L Pfaff, Todd S Macfarlan, and David J Katz in eLife. Published online January 27 2016 doi:10.7554/eLife.08848
Abstract
Maternally provided LSD1/KDM1A enables the maternal-to-zygotic transition and prevents defects that manifest postnatally
Somatic cell nuclear transfer has established that the oocyte contains maternal factors with epigenetic reprogramming capacity. Yet the identity and function of these maternal factors during the gamete to embryo transition remains poorly understood. In C. elegans, LSD1/KDM1A enables this transition by removing H3K4me2 and preventing the transgenerational inheritance of transcription patterns. Here we show that loss of maternal LSD1/KDM1A in mice results in embryonic arrest at the 1-2 cell stage, with arrested embryos failing to undergo the maternal-to-zygotic transition. This suggests that LSD1/KDM1A maternal reprogramming is conserved. Moreover, partial loss of maternal LSD1/KDM1A results in striking phenotypes weeks after fertilization; including perinatal lethality and abnormal behavior in surviving adults. These maternal effect hypomorphic phenotypes are associated with alterations in DNA methylation and expression at imprinted genes. These results establish a novel mammalian paradigm where defects in early epigenetic reprogramming can lead to defects that manifest later in development.
“Maternally provided LSD1/KDM1A enables the maternal-to-zygotic transition and prevents defects that manifest postnatally” by Jadiel A Wasson, Ashley K Simon, Dexter A Myrick, Gernot Wolf, Shawn Driscoll, Samuel L Pfaff, Todd S Macfarlan, and David J Katz in eLife. Published online January 27 2016 doi:10.7554/eLife.08848