Summary: Researchers produced the first cellular-resolution molecular map of human brain development in Down syndrome during the critical prenatal window. The study analyzed over 100,000 nuclei to resolve long-standing contradictions caused by inconsistent mouse models.
The findings reveal that Down syndrome disrupts the “tightly orchestrated sequence” of brain building: rather than cells simply dying off, the brain’s stem cells rush prematurely into neuron production. This depletion of the “progenitor pool” explains why individuals with Down syndrome often have smaller brain volumes and distinct cognitive profiles.
Key Facts
- The “Rush” to Differentiate: In typical development, progenitor (stem) cells divide to build a large pool before becoming neurons. In Down syndrome, these cells rush into differentiation too early, leading to an undersized “foundation” for the brain.
- The Neuron Skew: This premature shift creates a specific imbalance: an increase in upper-layer (IT) neurons (which connect the two brain hemispheres) and a reduction in deep-layer (CT) neurons (which connect the brain to the spinal cord for sensation and movement).
- Systems-Level Insights: Using multi-omics, researchers found that the disorder also alters cell metabolism and the way blood vessels (vasculature) interact with the nervous system, both of which likely accelerate the abnormal neuron production.
- Cross-Disorder Links: The molecular disruptions identified in Down syndrome overlap significantly with genetic risk factors for autism, epilepsy, and developmental delays, suggesting Down syndrome could serve as a “master model” for understanding neuropsychiatric conditions.
Source: UCLA
Scientists at UCLA have created one of the first cellular-resolution molecular maps detailing how Down syndrome alters human brain development before birth — a resource that resolves longstanding contradictions in the field and could lay the groundwork for future therapeutic strategies.
The study, published in Science, analyzed more than 100,000 nuclei from human prenatal neocortex samples collected across 26 pre-genotyped donors during gestational weeks 13 to 23 — the only window during which all the cortical neurons a person will carry for their entire life are generated.
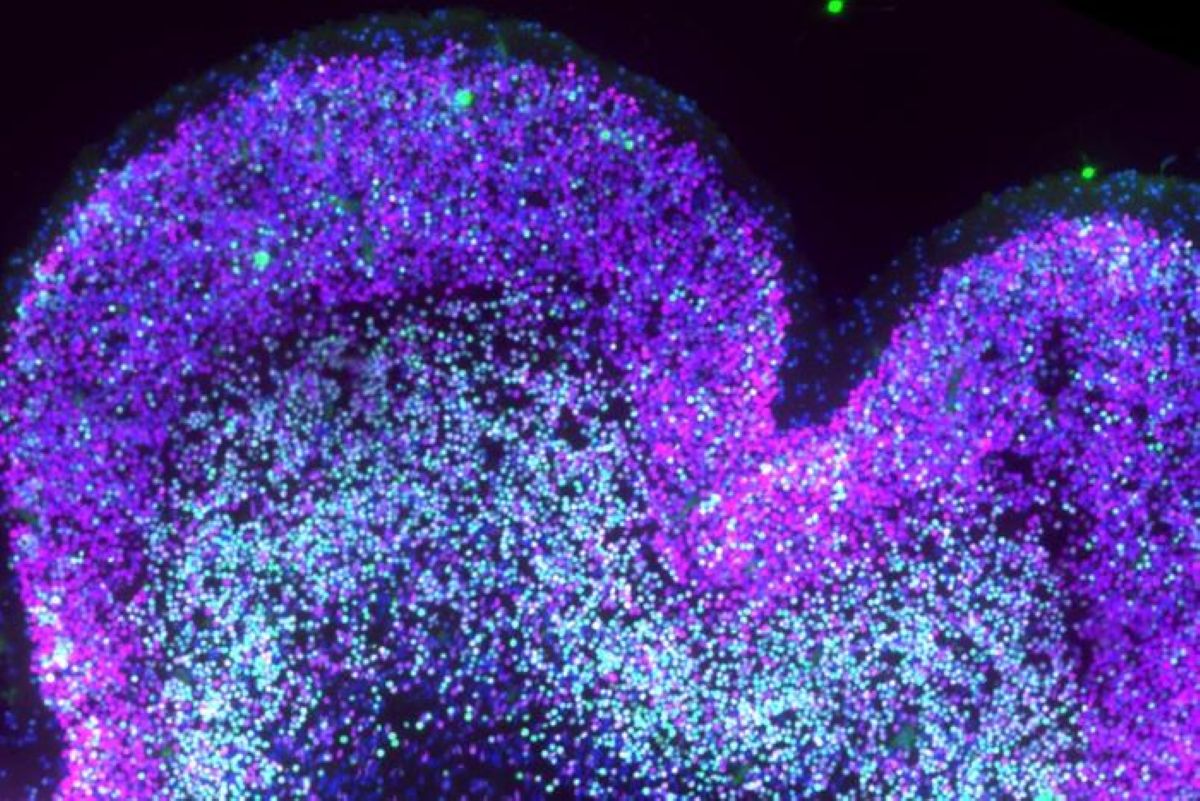
The findings suggest that Down syndrome disrupts the developmental sequence of that process, creating shifts that may help explain later differences in cognition, learning and sensory processing.
“There’s a new level of detail here that had never existed before,” said Luis de la Torre-Ubieta, senior author of the study and a member of the Eli and Edythe Broad Center of Regenerative Medicine and Stem Cell Research at UCLA. “For the first time, we can really try to understand systematically what’s going on in the developing brain of individuals with Down syndrome.”
Filling a critical gap
The Down syndrome research field has historically focused on two areas: the adult brain and the disorder’s connection to neurodegeneration. The link is striking — the vast majority of people with Down syndrome will develop Alzheimer’s disease, typically by their 60s.
What remained largely unexamined, despite clear indicators that Down syndrome is a developmental condition — such as smaller brain volumes detectable by MRI and cognitive differences apparent as early as 6 months to one year of age — was how the condition shapes the developing brain itself.
“No one had looked at the developing human brain in Down syndrome directly using single-cell genomics,” said de la Torre-Ubieta, an assistant professor of psychiatry and biobehavioral sciences. “Mouse models and in vitro models are important tools, but don’t really give you a gold standard of what’s happening in the human brain — and actually, they have led to different results and some confusion in the field.”
These inconsistencies are due in part to differences in how mice and human brains develop, and the fact that in vitro models don’t fully represent all the cell types and tissues present in the brain.
The new study, de la Torre-Ubieta said, can serve as that gold standard resource.
A disrupted developmental sequence and its impact on brain size
The development of the prenatal neocortex typically follows a tightly orchestrated sequence. Progenitor cells — the brain’s stem cells — must first divide repeatedly to expand their own pool, building up a sufficient base for all future neurons. Only then do they begin differentiating into neurons, starting with deep-layer cell types and progressing toward upper-layer cells in a carefully timed order.
In Down syndrome, that sequence appears to break down. The study found that progenitor cells appear to rush prematurely into neuron production, depleting their own pool and skewing the balance of neuron types generated. Specifically, the researchers observed a relative increase in upper-layer intratelencephalic neurons and a reduction in deep-layer corticothalamic neurons.
Those two cell populations play fundamentally different roles: CT neurons project outward from the cortex — connecting to brain structures and to the spinal cord to govern sensation and movement; IT neurons wire within the cortex, connecting the two hemispheres and contributing to information processing. This finding offers a new hypothesis for how early developmental changes might contribute to the cognitive profile of the condition.
The finding also offers a new answer to a longstanding question in the field: Why do people with Down syndrome tend to have smaller brains? Earlier theories centered on elevated rates of cell death. The current study found less evidence of widespread neuronal death and instead points to the depletion of the progenitor pool.
A systems-level view of a systems-level disorder
The study employed paired single-nucleus multi-omics, a technology that measures both gene expression and chromatin accessibility in the same individual cell. Chromatin accessibility reveals which regions of the genome are open and active — the enhancers and promoters that regulate gene expression — offering a layer of information beyond which genes are simply switched on or off.
By combining these two readouts, the researchers were able to reconstruct not just a snapshot of which cells are present, but the regulatory programs that guide cell fate — and how those programs are disrupted in Down syndrome. Systems-level approaches also led them to uncover alterations in cell metabolism and changes in how the vasculature interacts with the developing nervous system, both of which could speed up neuron production.
Implications beyond Down syndrome
The study’s significance extends beyond Down syndrome. The researchers specifically tested for overlap between the molecular disruptions they identified and the genetic risk signatures associated with other neurodevelopmental and neuropsychiatric conditions, including autism, epilepsy and developmental delay. They found substantial convergence, particularly in the gene-regulatory networks governing the specification of IT versus CT neurons.
“Down syndrome could be a model to understand intellectual disability and neuropsychiatric disorders more broadly,” de la Torre-Ubieta said. “Also to uncover the shared biology underlying these conditions — because the mechanisms are often still unknown.”
Two papers, one story
The publication coincides with a companion paper from researchers at the University of Wisconsin-Madison, appearing in the same issue of Science. While the UCLA study focuses on the prenatal period, the Wisconsin team examined the postnatal brain, studying Down syndrome between approximately one and five years of age.
When the two groups shared preliminary findings, they discovered striking parallels: many of the changes identified prenatally by the UCLA team appear to persist into early childhood.
Together, the two papers provide a continuous molecular view of Down syndrome brain development from mid-gestation through infancy — a resource that did not previously exist and that the researchers expect will serve as a reference for their field for years to come.
A foundation for future therapies
While the researchers are careful to emphasize that the findings do not point to a near-term clinical application, the study provides the clearest picture yet of the cellular and molecular events that distinguish the Down syndrome brain during development, and a framework for identifying future therapeutic targets.
“We are finding targets that could be actionable down the line if you generate drugs for specific pathways,” de la Torre-Ubieta said. “And you could conceive of a gene therapy based on that, to suppress the expression of particular drivers and restore development closer to its normal course.”
UCLA authors Celine K. Vuong and Alexis Weber led this work, along with Patrick Seong, Yu-Jen Chen, Jordan Peyer, Shahab Younesi, Angelo Salinda, Daniel Gomez, Gabriella Rivas, Abril Morales, Beck Shafie, Pan Zhang, Susanne Nichterwitz, Le Qi, Nolan T. Fernandez, Emily Friedman, Daniel H. Geschwind and William E. Lowry. Nana Matoba, Michael I. Love, Michael J. Gandal and Jason L. Stein contributed to this study.
Funding: The research was supported by the National Institute of Child Health and Human Development, the National Institute of Mental Health, the UCLA Broad Stem Cell Research Center, including a Rose Hills Foundation Innovator Grant and a post-doctoral training grant, the UCLA Health Jonsson Comprehensive Cancer Center and UCLA Broad Stem Cell Research Center Ablon Scholars Program, the California Institute for Regenerative Medicine and the National Institutes of Health Biomedical Big Data Training Program.
Key Questions Answered:
A: Mouse brains develop very differently from human brains, which has led to “scientific noise” and confusion. This study used actual human prenatal samples during the exact 10-week window (gestational weeks 13–23) when every cortical neuron a human will ever have is created. It is now considered the “gold standard” reference for the field.
A: While this study focused on early development, it provides the “starting line” data. By understanding how the brain is built differently from the beginning, scientists can better understand why those same neural networks are more vulnerable to neurodegeneration and Alzheimer’s later in life.
A: The researchers are cautious but optimistic. By identifying the specific “drivers” that cause stem cells to rush into neuron production, they have identified potential targets for future gene therapies or drugs that could help restore the developmental sequence to a more normal course.
Editorial Notes:
- This article was edited by a Neuroscience News editor.
- Journal paper reviewed in full.
- Additional context added by our staff.
About this neurodevelopment and Down syndrome research news
Author: Ani Vahradyan
Source: UCLA
Contact: Ani Vahradyan – UCLA
Image: The image is credited to de la Torre-Ubieta Lab
Original Research: Closed access.
“A single-cell multiomic analysis identifies molecular and gene-regulatory mechanisms dysregulated in developing Down syndrome neocortex” by Celine K. Vuong, Alexis Weber, Patrick Seong, Nana Matoba, Yu-Jen Chen, Jordan Peyer, Shahab Younesi, Angelo Salinda, Daniel Gomez, Gabriella Rivas, Abril Morales, Beck Shafie, Pan Zhang, Susanne Nichterwitz, Le Qi, Nolan T. Fernandez, Emily Friedman, Michael I. Love, Michael J. Gandal, Daniel H. Geschwind, William E. Lowry, Jason L. Stein, and Luis de la Torre-Ubieta. Science
DOI:10.1126/science.aea1259
Abstract
A single-cell multiomic analysis identifies molecular and gene-regulatory mechanisms dysregulated in developing Down syndrome neocortex
INTRODUCTION
Down syndrome (DS), caused by triplication of human chromosome 21 (Ts21), is the most common genetic cause of intellectual disability. Individuals with DS show deficits in learning, memory, and attention; delayed language and motor development; and atypical sensory processing.
The early emergence of structural abnormalities in the neocortex and of cognitive defects suggests that corticogenesis is disrupted in DS. Yet the molecular and cellular mechanisms leading to changes in the developing DS brain remain to be elucidated.
RATIONALE
Precise temporal and molecular processes driving cell type–specific gene expression programs govern the development of the human brain, a process not yet completely captured by either in vitro or animal models of DS. To address this, we leveraged single-nucleus multiomics (gene expression plus chromatin accessibility) in developing control (Ctrl) and Ts21 neocortex during the period of peak neurogenesis to characterize the cellular and molecular processes and the gene-regulatory mechanisms underpinning DS.
RESULTS
We profiled neocortex from a cohort of 26 Ctrl and Ts21 donors from gestation weeks (GW) 13 to 23 using paired single-nucleus multiomics to capture gene expression and regions of open chromatin across 113,000 nuclei. We uncovered altered cell composition in the Ts21 neocortex, including a reduction in progenitors and misspecification of excitatory neurons.
The neurogenic timeline was accelerated in Ts21, with increased commitment of newborn neurons to the upper layer intratelencephalic (IT) neuron fate at the expense of deep layer corticothalamic (CT) neurons. We then systematically assessed gene expression and coexpression networks altered in Ts21 to identify molecular pathways underlying these cellular changes.
This revealed widespread gene dysregulation encompassing proliferative and metabolic pathways involved in maintaining the progenitor pool, as well as aberrant expression of pro-IT gene programs in newborn and deep layer neurons.
Altered cell composition, timing of neurogenesis, and molecular programs were recapitulated by using primary human neural progenitor cells derived from Ctrl and Ts21 donors. We used gene expression to examine intercellular interactions, finding proneurogenic changes in the neurovascular niche and early evidence of microglial activation.
We then combined paired gene expression and chromatin accessibility to identify gene-regulatory networks and the transcription factors controlling gene programs altered in Ts21. Among these we identified a human chromosome 21 (HSA21)–encoded TF, BACH1, as an activator of pro-IT programs.
We leveraged new knowledge of these gene expression and regulatory changes to understand the molecular mechanisms shared between DS and other neurodevelopmental and psychiatric disorders with cognitive impairment, identifying deep layer neurons and specification programs as a point of shared vulnerability.
CONCLUSION
Our study reveals the neurodevelopmental changes occurring in the Ts21 neocortex and defines the gene-regulatory mechanisms driving them. Our findings offer new insight into the earliest steps leading to DS and provide a foundation for future therapeutic targets.