Summary: Researchers have identified cognitive subgroups related to genetic differences in Alzheimer’s patients. The findings, researchers say, could open the door for more personalized treatments of the neurodegenerative disease.
Source: UW Medicine.
Researchers studying Alzheimer’s disease have created an approach to classify patients with Alzheimer’s disease, a finding that may open the door for personalized treatments.
“Alzheimer’s, like breast cancer, is not one disease,” said lead author Shubhabrata Mukherjee, research assistant professor in general internal medicine at the University of Washington School of Medicine. “I think a good drug might fail in a clinical trial because not all the subjects have the same kind of Alzheimer’s.
This study, published in the recent issue of Molecular Psychiatry, involves 19 researchers from several institutions, including Boston University School of Medicine, the VA Puget Sound Health Care System and Indiana University School of Medicine.
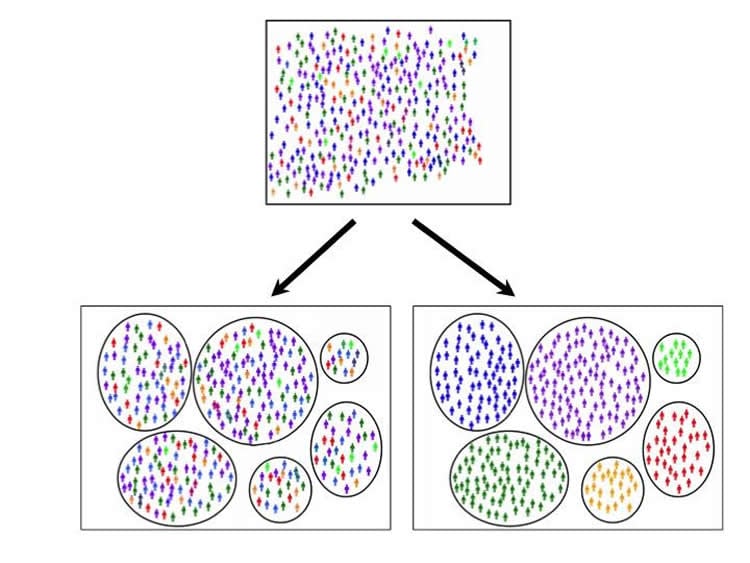
The researchers put 4,050 people with late-onset Alzheimer’s disease into six groups based on their cognitive functioning at the time of diagnosis and then used genetic data to find biological differences across these groups.
“The implications are exciting,” said corresponding author Paul Crane, professor of general internal medicine at the University of Washington School of Medicine. “We have found substantial biological differences among cognitively defined subgroups of Alzheimer’s patients.”
Identification of cognitive subgroups related to genetic differences is an important step toward developing a precision medicine approach for Alzheimer’s disease.
The participants received cognitive scores in four domains: memory, executive functioning, language, and visuospatial functioning.
The largest group (39%) had scores in all four domains that were fairly close to each other. The next largest group (27%) had memory scores substantially lower than their other scores. Smaller groups had language scores substantially lower than their other scores (13%), visuospatial functioning scores substantially lower than their other scores (12%), and executive functioning scores substantially lower than their other scores (3%). There were 6% who had two domains that were substantially lower than their other scores.
The participants came from five studies, and it took more than two years to standardize the neuropsychological test scores across all the studies in order to detect meaningful patterns. The mean age was 80, 92 percent self-reported white race, and 61 percent were female.
The investigators used genome-wide genetic data to find out if the subgroups are biologically distinct.
Investigators found 33 single nucleotide polymorphisms (SNPs) – specific locations throughout the genome – where the genetic association was very strong for one of the subgroups. These genetic relationships were stronger than the strongest effects found by an earlier and much larger international consortium study where Alzheimer’s disease was treated as a single homogeneous condition.
Several years ago, the International Genomics of Alzheimer’s Project Consortium published the largest genome-wide association study of Alzheimer’s disease and found about 20 SNPs associated with Alzheimer’s disease risk.
This study found 33 additional SNPs with even stronger relationships with a single subgroup.
The study also found a particularly strong relationship between a particular variant of the APOE gene and risk for the memory subgroup. The APOE e4 allele is a very strong risk factor for developing Alzheimer’s disease for people with European ancestry, and it also appears to influence which cognitive subtype of Alzheimer’s a person is likely to develop.
People can currently find out if they have an APOE e4 allele with direct-to-consumer testing; however, the researchers note that many people with an APOE e4 allele never develop Alzheimer’s disease, and many who don’t carry any known genetic risk factor nevertheless end up with the condition.
While world leaders want to find a cure for Alzheimer’s by 2025, so far no one has been able to develop an effective treatment let alone a cure. But this study suggests that thinking of Alzheimer’s disease as six distinct conditions may provide a way forward.
“This study is not the end, it’s a start,” said Mukherjee.
Currently, 5.7 million people in the United States are living with the disease and that is expected to grow to 14 million by 2050, according to the Alzheimer’s Association.
The research team also included investigators from the Department of Biostatistics at University of Kentucky, Department of Epidemiology at John Hopkins Bloomberg School of Public Health, departments of neurology, human genetics and psychiatry at the University of Pittsburgh, Center for Translational & Computational Neuroimmunology at Columbia University Medical Center, Rush Alzheimer’s Disease Center at Rush University Medical Center in Chicago, and Kaiser Permanente Washington Health Research Institute in Seattle.
Funding: This research was supported by the Nancy and Buster Alvord Endowment, Department of Veterans Affairs Research Funds, Alzheimer’s Disease Neuroimaging Initiative, National Institute of Aging, National Institute of Biomedical Imaging and Bioengineering, Alzheimer’s Association, and many more.
The authors report no financial or other conflicts of interest.
Source: Bobbi Nodell – UW Medicine
Publisher: Organized by NeuroscienceNews.com.
Image Source: NeuroscienceNews.com image is credited to UW Medicine.
Original Research: Abstract for “Genetic data and cognitively defined late-onset Alzheimer’s disease subgroups” Shubhabrata Mukherjee, Jesse Mez, Emily H. Trittschuh, Andrew J. Saykin, Laura E. Gibbons, David W. Fardo, Madeline Wessels, Julianna Bauman, Mackenzie Moore, Seo-Eun Choi, Alden L. Gross, Joanne Rich, Diana K. N. Louden, R. Elizabeth Sanders, Thomas J. Grabowski, Thomas D. Bird, Susan M. McCurry, Beth E. Snitz, M. Ilyas Kamboh, Oscar L. Lopez, Philip L. De Jager, David A. Bennett, C. Dirk Keene, Eric B. Larson, EPAD Study Group, Investigators from ACT, Investigators from ROS, Investigators from MAP, Investigators from ADNI, Investigators from the University of Pittsburgh ADRC & Paul K. Crane in Molecular Psychiatry. Published December 4 2018.
doi:10.1038/s41380-018-0298-8
[cbtabs][cbtab title=”MLA”]UW Medicine”Alzheimer’s Patients Classified in Six Subgroups.” NeuroscienceNews. NeuroscienceNews, 5 December 2018.
<https://neurosciencenews.com/alzheimers-subgroups-10308/>.[/cbtab][cbtab title=”APA”]UW Medicine(2018, December 5). Alzheimer’s Patients Classified in Six Subgroups. NeuroscienceNews. Retrieved December 5, 2018 from https://neurosciencenews.com/alzheimers-subgroups-10308/[/cbtab][cbtab title=”Chicago”]UW Medicine”Alzheimer’s Patients Classified in Six Subgroups.” https://neurosciencenews.com/alzheimers-subgroups-10308/ (accessed December 5, 2018).[/cbtab][/cbtabs]
Abstract
Genetic data and cognitively defined late-onset Alzheimer’s disease subgroups
Categorizing people with late-onset Alzheimer’s disease into biologically coherent subgroups is important for personalized medicine. We evaluated data from five studies (total n = 4050, of whom 2431 had genome-wide single-nucleotide polymorphism (SNP) data). We assigned people to cognitively defined subgroups on the basis of relative performance in memory, executive functioning, visuospatial functioning, and language at the time of Alzheimer’s disease diagnosis. We compared genotype frequencies for each subgroup to those from cognitively normal elderly controls. We focused on APOE and on SNPs with p < 10−5 and odds ratios more extreme than those previously reported for Alzheimer’s disease (<0.77 or >1.30). There was substantial variation across studies in the proportions of people in each subgroup. In each study, higher proportions of people with isolated substantial relative memory impairment had ≥1 APOE ε4 allele than any other subgroup (overall p = 1.5 × 10−27). Across subgroups, there were 33 novel suggestive loci across the genome with p < 10−5 and an extreme OR compared to controls, of which none had statistical evidence of heterogeneity and 30 had ORs in the same direction across all datasets. These data support the biological coherence of cognitively defined subgroups and nominate novel genetic loci.