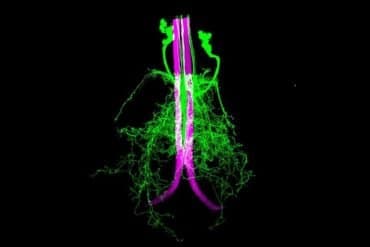
Summary: Computational neurosciences have created a virtual version of Purkinje cells in the brain’s cerebellum. The virtual cells may shed light on how Purkinje cells work.
Source: OIST.
There are no crystal balls in science. For researchers, making predictions about the outcome of an experiment requires close observation of available data. But in neurobiology, making accurate observations is hampered by the difficulty of measuring the cells that control everything from a finger click to philosophy..
There is a workaround, however: modeling. Simulating the world based on experimental observation allows for accurate predictions, and with sophisticated programming, sophisticated models are possible. That’s the case with a computational model of a crucial brain cell, which has just had a thorough upgrade by a scientist at the Okinawa Institute of Science and Technology Graduate University (OIST). Their study was published in Cell Reports.
This latest model created a virtual version of brain cells known as Purkinje cells. These large nerve cells are found in the brain’s cerebellum, where they help control the accurate movement of the body. Given this crucial function, it’s no surprise that there is a demand for greater understanding about how they work.
To this end, Belgian neuroscientist and head of OIST’s Computational Neuroscience Unit, Prof. Erik De Schutter proposed a model to explain its function in 1994, but knew there was room for improvement: “new data would arrive, and we learned there were aspects of the model that couldn’t be correct”, said Prof. De Schutter. There was a need for an updated model, but it would be a big time investment and have to set a new standard for the field. No small task.
This is where Dr. Yunliang Zang entered the picture. Coming from a background of modeling cardiac function, Zang found his own heart drawn to the research projects in neuroscience at OIST. “In cardiac modeling, we have to achieve an extensively validated model,” said Dr. Zang, “But in computational neuroscience, things are more sloppy. I wanted to do better.”
Dr. Zang used this determination to create a model based on experimental data that shows how Purkinje cells can be taught by neural structures, called climbing fibers, to perform their functions more efficiently. Doing that, however, required building a virtual version of the neuron.
Making this virtual cell is a little like trying to build a cellphone: you need to start with the individual chips and circuitry and put them together in the right way. When it comes to the Purkinje cells, the “chips” are the individual neuronal structures. Previous models were based on experimental readings from the whole neuron, but Dr. Zang’s breakthrough lay in building his model from exact recordings from their constituent parts. The result was like assembling the phone chips to make a functional whole.
But for a model to stand on its feet, it needs to accommodate all the possible outcomes of a variety of situations and give the correct answer every time. Getting to that point meant a lot of reverse engineering using the results from existing studies. “I had to beg scientists from all over the world to share their study data with me,” said Dr. Zang, “getting as much as possible was crucial to get the most complete picture”.
With large volumes of data, there are large volumes of problems. Experimental data from one research group would sometimes contradict that from others. Dr. Zang faced the daunting prospect of reconciling these conflicting observations in the model. “I spent two and a half years going back and forth, making sure the model could replicate experimental observations, and I found that the key factor to these differences was the variation in experimental conditions.” The result is a more robust and reliable model that could be used to clearly explain the inner workings of the neurons.
Imagine you want to touch your cellphone: It’s easy for your finger to tap the phone generally, but more of a feat to accurately press the specific icon that takes you to your latest twitter meme. Your cerebellum acts in real time to help you zero in on a target. It is thought to achieve this via neural structures called climbing fibers, which can “teach” other neurons in the cerebellum the correct ways to achieve accurate movement.
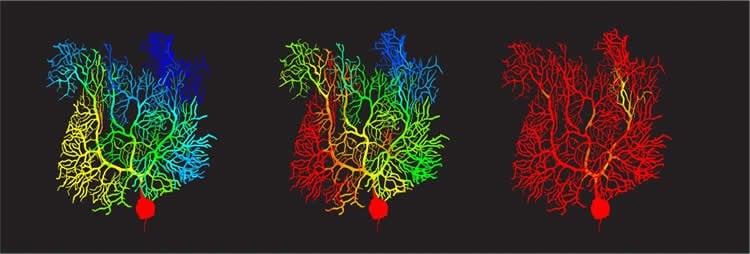
Previously, these climbing fiber signals were thought to be binary in their effects, like a guide telling you to move or not to move your hand). But Zang’s study shows that climbing fibers can teach Purkinje cells to fire with analogue information, such as pressing a specific button. Instead of the Purkinje cells responding to the signals with an all-or-nothing signal to the body, Zang’s model demonstrates they are capable of in-between signals, allowing for the nuance that makes fine targeting possible.
The effect comes down to something called a potassium current, which acts like a brake on signals travelling in the Purkinje cell dendritic trees. Using his simulations, Zang found that varying braking levels by voltage-dependent potassium current achieved the learning effect he was looking for in the cells.
This is just the tip of the iceberg for this model, which promises to serve as a new, more stable foundation for all researchers in the field. “I’m extremely happy. This new model will have a very long shelf life,” said Prof. De Schutter, “and it’s going to be a new standard for the field”.
Source: Kaoru Natori – OIST
Publisher: Organized by NeuroscienceNews.com.
Image Source: NeuroscienceNews.com image is credited to OIST.
Original Research: Open access research for “Voltage- and Branch-Specific Climbing Fiber Responses in Purkinje Cells” by Yunliang Zang, Stéphane Dieudonné, Erik De Schutter in Cell Reports. Published August 7 2018.
doi:10.1016/j.celrep.2018.07.011
[cbtabs][cbtab title=”MLA”]OIST”Upgrade for Virtual Brain Cell.” NeuroscienceNews. NeuroscienceNews, 7 August 2018.
<https://neurosciencenews.com/virtual-brain-cell-9666/>.[/cbtab][cbtab title=”APA”]OIST(2018, August 7). Upgrade for Virtual Brain Cell. NeuroscienceNews. Retrieved August 7, 2018 from https://neurosciencenews.com/virtual-brain-cell-9666/[/cbtab][cbtab title=”Chicago”]OIST”Upgrade for Virtual Brain Cell.” https://neurosciencenews.com/virtual-brain-cell-9666/ (accessed August 7, 2018).[/cbtab][/cbtabs]
Abstract
Voltage- and Branch-Specific Climbing Fiber Responses in Purkinje Cells
Climbing fibers (CFs) provide instructive signals driving cerebellar learning, but mechanisms causing the variable CF responses in Purkinje cells (PCs) are not fully understood. Using a new experimentally validated PC model, we unveil the ionic mechanisms underlying CF-evoked distinct spike waveforms on different parts of the PC. We demonstrate that voltage can gate both the amplitude and the spatial range of CF-evoked Ca2+ influx by the availability of K+ currents. This makes the energy consumed during a complex spike (CS) also voltage dependent. PC dendrites exhibit inhomogeneous excitability with individual branches as computational units for CF input. The variability of somatic CSs can be explained by voltage state, CF activation phase, and instantaneous CF firing rate. Concurrent clustered synaptic inputs affect CSs by modulating dendritic responses in a spatially precise way. The voltage- and branch-specific CF responses can increase dendritic computational capacity and enable PCs to actively integrate CF signals.