Summary: Rare structural genetic variants could play a role in the development of schizophrenia.
Source: UNC Health Care
Most research about the genetics of schizophrenia has sought to understand the role that genes play in the development and heritability of schizophrenia. Many discoveries have been made, but there have been many missing pieces. Now, UNC School of Medicine scientists have conducted the largest-ever whole genome sequencing study of schizophrenia to provide a more complete picture of the role the human genome plays in this disease.
Published in Nature Communications, the study co-led by senior author Jin Szatkiewicz, PhD, associate professor in the UNC Department of Genetics, suggests that rare structural genetic variants could play a role in schizophrenia.
“Our results suggest that ultra-rare structural variants that affect the boundaries of a specific genome structure increase risk for schizophrenia,” Szatkiewicz said. “Alterations in these boundaries may lead to dysregulation of gene expression, and we think future mechanistic studies could determine the precise functional effects these variants have on biology.”
Previous studies on the genetics of schizophrenia have primarily involved using common genetic variations known as SNPs (alterations in common genetic sequences and each affecting a single nucleotide), rare variations in the part of DNA that provide instructions for making proteins, or very large structural variations (alterations affecting a few hundred thousands of nucleotides). These studies give snapshots of the genome, leaving a large portion of the genome a mystery, as it potentially relates to schizophrenia.
In the Nature Communications study, Szatkiewicz and colleagues examined the entire genome, using a method called whole genome sequencing (WGS). The primary reason WGS hasn’t been more widely used is that it is very expensive. For this study, an international collaboration pooled funding from National Institute of Mental Health grants and matching funds from Sweden’s SciLife Labs to conduct deep whole genome sequencing on 1,165 people with schizophrenia and 1,000 controls – the largest known WGS study of schizophrenia ever.
As a result, new discoveries were made. Previously undetectable mutations in DNA were found that scientists had never seen before in schizophrenia.
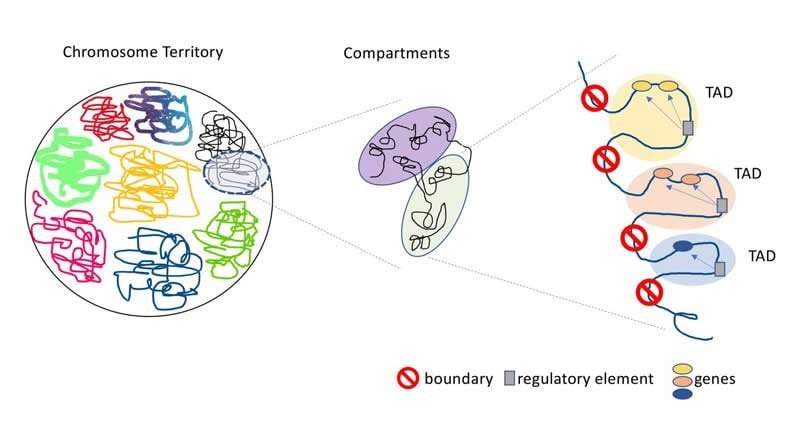
In particular, this study highlighted the role that a three-dimensional genome structure known as topologically associated domains (TADs) could play in the development of schizophrenia. TADs are distinct regions of the genome with strict boundaries between them that keep the domains from interacting with genetic material in neighboring TADs. Shifting or breaking these boundaries allows interactions between genes and regulatory elements that normally would not interact.
When these interactions occur, gene expression may be changed in undesirable ways that could result in congenital defects, formation of cancers, and developmental disorders. This study found that extremely rare structural variants affecting TAD boundaries in the brain occur significantly more often in people with schizophrenia than in those without it. Structural variants are large mutations that may involve missing or duplicated genetic sequences, or sequences that are not in the typical genome. This finding suggests that misplaced or missing TAD boundaries may also contribute to the development of schizophrenia. This study was the first to discover the connection between anomalies in TADs and the development of schizophrenia.
This work has highlighted TADs-affecting structural variants as prime candidates for future mechanistic studies of the biology of schizophrenia.
“A possible future investigation would be to work with patient-derived cells with these TADs-affecting mutations and figure out what exactly happened at the molecular level,” said Szatkiewicz, an adjunct assistant professor of psychiatry at UNC. “In the future, we could use this information about the TAD effects to help develop drugs or precision medicine treatments that could repair disrupted TADs or affected gene expressions which may improve patient outcomes.”
This study will be combined with other WGS studies in order to increase the sample size to further confirm these results. This research will also help the scientific community build on the unfolding genetic mysteries of schizophrenia.
Patrick Sullivan, MD, the Yeargen Distinguished Professor of Psychiatry and Genetics at the UNC School of Medicine and Director of the Center for Psychiatric Genomics, is co-senior author. Co-first authors from UNC are Matthew Halvorsen, PhD, Ruth Huh, PhD, Jia Wen, PhD, from the UNC Department of Genetics. Other UNC authors are Paola Giusto-Rodriguez, PhD, NaEshia Ancalade, Martilias Farrell, PhD, James Crowley, PhD, and Yun Li, PhD.
Funding: This research was a collaboration between researchers at UNC-Chapel Hill, Lund University, Chalmers University of Technology, the Karolinska Institutet, and Uppsala University.
About this genetics research article
Source:
UNC Health Care
Media Contacts:
Mark Derewicz – UNC Health Care
Image Source:
The image is credited to Jin Szatkiewicz, PhD, UNC School of Medicine.
Original Research: Open access
“Increased burden of ultra-rare structural variants localizing to boundaries of topologically associated domains in schizophrenia”. by Matthew Halvorsen, Ruth Huh, Nikolay Oskolkov, Jia Wen, Sergiu Netotea, Paola Giusti-Rodriguez, Robert Karlsson, Julien Bryois, Björn Nystedt, Adam Ameur, Anna K. Kähler, NaEshia Ancalade, Martilias Farrell, James J. Crowley, Yun Li, Patrik K. E. Magnusson, Ulf Gyllensten, Christina M. Hultman, Patrick F. Sullivan & Jin P. Szatkiewicz.
Nature Communications doi:10.1038/s41467-020-15707-w.
Abstract
Increased burden of ultra-rare structural variants localizing to boundaries of topologically associated domains in schizophrenia
Despite considerable progress in schizophrenia genetics, most findings have been for large rare structural variants and common variants in well-imputed regions with few genes implicated from exome sequencing. Whole genome sequencing (WGS) can potentially provide a more complete enumeration of etiological genetic variation apart from the exome and regions of high linkage disequilibrium. We analyze high-coverage WGS data from 1162 Swedish schizophrenia cases and 936 ancestry-matched population controls. Our main objective is to evaluate the contribution to schizophrenia etiology from a variety of genetic variants accessible to WGS but not by previous technologies. Our results suggest that ultra-rare structural variants that affect the boundaries of topologically associated domains (TADs) increase risk for schizophrenia. Alterations in TAD boundaries may lead to dysregulation of gene expression. Future mechanistic studies will be needed to determine the precise functional effects of these variants on biology.
Feel Free To Share This Schizophrenia News.






