A new imaging tool developed by Boston scientists could do for the brain what the telescope did for space exploration. In the first demonstration of how the technology works, published July 30 in the journal Cell, the researchers look inside the brain of an adult mouse at a scale previously unachievable, generating images at a nanoscale resolution. The inventors’ long-term goal is to make the resource available to the scientific community in the form of a national brain observatory.
“I’m a strong believer in bottom up-science, which is a way of saying that I would prefer to generate a hypothesis from the data and test it,” says senior study author Jeff Lichtman, of Harvard University. “For people who are imagers, being able to see all of these details is wonderful and we’re getting an opportunity to peer into something that has remained somewhat intractable for so long. It’s about time we did this, and it is what people should be doing about things we don’t understand.”
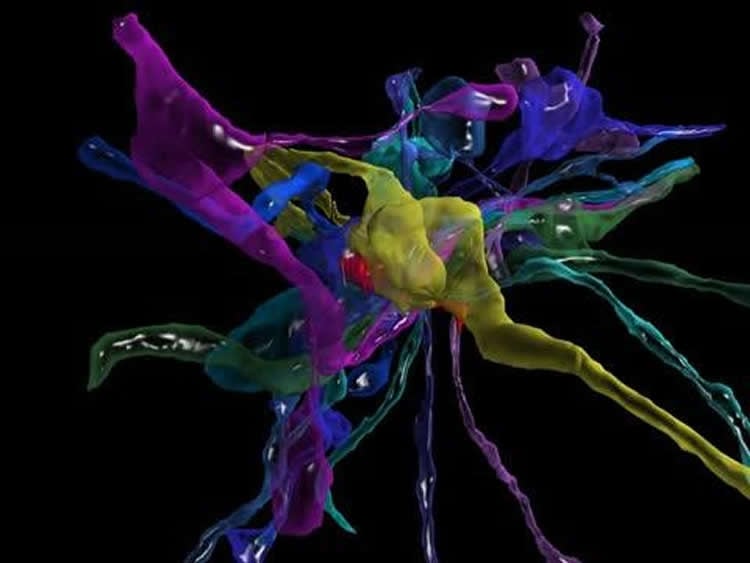
The researchers have begun the process of mining their imaging data by looking first at an area of the brain that receives sensory information from mouse whiskers, which help the animals orient themselves and are even more sensitive than human fingertips. The scientists used a program called VAST, developed by co-author Daniel Berger of Harvard and the Massachusetts Institute of Technology, to assign different colors and piece apart each individual “object” (e.g., neuron, glial cell, blood vessel cell, etc.).
“The complexity of the brain is much more than what we had ever imagined,” says study first author Narayanan “Bobby” Kasthuri, of the Boston University School of Medicine. “We had this clean idea of how there’s a really nice order to how neurons connect with each other, but if you actually look at the material it’s not like that. The connections are so messy that it’s hard to imagine a plan to it, but we checked and there’s clearly a pattern that cannot be explained by randomness.”
The researchers see great potential in the tool’s ability to answer questions about what a neurological disorder actually looks like in the brain, as well as what makes the human brain different from other animals and different between individuals. Who we become is very much a product of the connections our neurons make in response to various life experiences. To be able to compare the physical neuron-to-neuron connections in an infant, a mathematical genius, and someone with schizophrenia would be a leap in our understanding of how our brains shape who we are (or vice versa).
The cost and data storage demands for this type of research are still high, but the researchers expect expenses to drop over time (as has been the case with genome sequencing). To facilitate data sharing, the scientists are now partnering with Argonne National Laboratory with the hopes of creating a national brain laboratory that neuroscientists around the world can access within the next few years.
“It’s bittersweet that there are many scientists who think this is a total waste of time as well as a big investment in money and effort that could be better spent answering questions that are more proximal,” Lichtman says. “As long as data is showing you things that are unexpected, then you’re definitely doing the right thing. And we are certainly far from being out of the surprise element. There’s never a time when we look at this data that we don’t see something that we’ve never seen before.”
Funding: This work was supported by the NIH/NINDS, Conte, the MURI Army Research Office, the NSF, DARPA, the Human Frontier Science Program, the JHU Applied Physics Laboratory, the Research Program for Applied Neuroscience, the Howard Hughes Medical Institute, Nvidia, Intel, and Google.
Source: Joseph Caputo – Cell Press
Image Credit: The image is credited to the Kasthuri et al./Cell 2015
Original Research: Abstract for “Saturated Reconstruction of a Volume of Neocortex ” by Narayanan Kasthuri, Kenneth Jeffrey Hayworth, Daniel Raimund Berger, Richard Lee Schalek, José Angel Conchello, Seymour Knowles-Barley, Dongil Lee, Amelio Vázquez-Reina, Verena Kaynig, Thouis Raymond Jones, Mike Roberts, Josh Lyskowski Morgan, Juan Carlos Tapia, H. Sebastian Seung, William Gray Roncal, Joshua Tzvi Vogelstein, Randal Burns, Daniel Lewis Sussman, Carey Eldin Priebe, Hanspeter Pfister, and Jeff William Lichtman in Cell. Published online June 1 2015 doi:10.1016/j.cell.2015.06.054
Abstract
Saturated Reconstruction of a Volume of Neocortex
Highlights
•Tape-based pipeline for electron microscopic reconstruction of brain tissue
•Annotated database of 1,700 synapses from a saturated reconstruction of cortex
•Excitatory axon proximity to dendritic spines not sufficient to predict synapses
Summary
We describe automated technologies to probe the structure of neural tissue at nanometer resolution and use them to generate a saturated reconstruction of a sub-volume of mouse neocortex in which all cellular objects (axons, dendrites, and glia) and many sub-cellular components (synapses, synaptic vesicles, spines, spine apparati, postsynaptic densities, and mitochondria) are rendered and itemized in a database. We explore these data to study physical properties of brain tissue. For example, by tracing the trajectories of all excitatory axons and noting their juxtapositions, both synaptic and non-synaptic, with every dendritic spine we refute the idea that physical proximity is sufficient to predict synaptic connectivity (the so-called Peters’ rule). This online minable database provides general access to the intrinsic complexity of the neocortex and enables further data-driven inquiries.
“Saturated Reconstruction of a Volume of Neocortex ” by Narayanan Kasthuri, Kenneth Jeffrey Hayworth, Daniel Raimund Berger, Richard Lee Schalek, José Angel Conchello, Seymour Knowles-Barley, Dongil Lee, Amelio Vázquez-Reina, Verena Kaynig, Thouis Raymond Jones, Mike Roberts, Josh Lyskowski Morgan, Juan Carlos Tapia, H. Sebastian Seung, William Gray Roncal, Joshua Tzvi Vogelstein, Randal Burns, Daniel Lewis Sussman, Carey Eldin Priebe, Hanspeter Pfister, and Jeff William Lichtman in Cell. Published online June 1 2015 doi:10.1016/j.cell.2015.06.054