A new study by Valerie Hu, Ph.D., professor of biochemistry and molecular medicine at the George Washington University (GW) School of Medicine and Health Sciences (SMHS), reports that RORA, a novel candidate gene for autism discovered by her group in a 2010 study, regulates a large number of other genes associated with autism.
“We are focusing on this gene, in part, because this gene can act as a master regulator of other genes,” said Hu, whose study was published in the journal Molecular Autism. “Called nuclear hormone receptors, they are capable of activating or suppressing other genes in the genome. The question was which specific genes are regulated by RORA.”
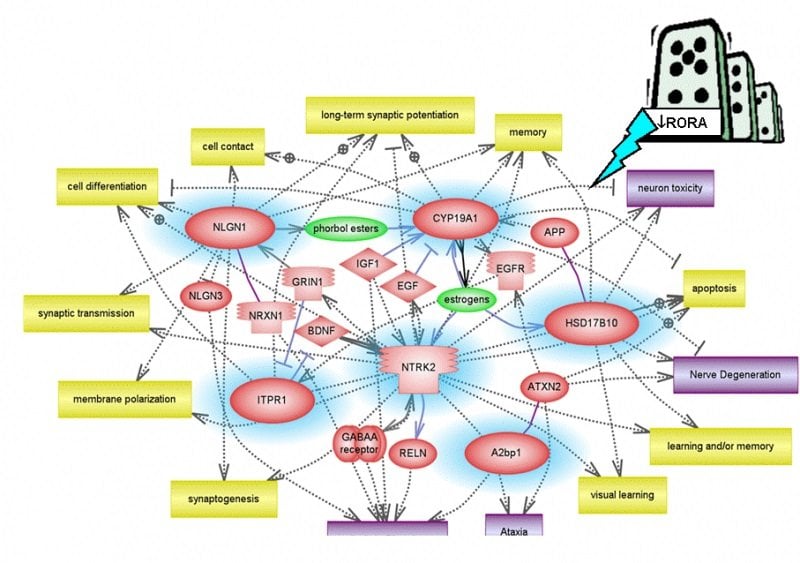
Hu and co-author, Tewarit Sarachana, Ph.D., a former doctoral student in the molecular medicine doctoral program at SMHS, found that RORA encodes a protein that can regulate the expression of more than 2,500 other genes. Of these 2,500 genes, many are known to be involved in neuronal development and functions, and 426 of RORA’s gene targets are already listed in AutismKB, a database of known autism candidate genes.

To identify genes regulated by RORA, Hu and Sarachana used chromatin immunoprecipitation (ChIP) with an anti-RORA antibody followed by whole-genome promoter array (chip) analysis. This genome-wide ChIP-on-chip analysis of target genes of RORA, as well as additional methods of validation, confirmed that RORA transcriptionally regulates the genes A2BP1, CYP19A1, HSD17B10, ITPR1, NLGN1, and NTRK2, such that when RORA levels are cut in half, all six genes also go down in their expression. The expression levels of these six genes are also reduced in RORA-deficient postmortem brain tissues from individuals with autism relative to that of age-matched unaffected controls.
“We see it as a domino effect, where RORA is a particularly shaky domino,” said Hu. “If knocked over, it can also knock down a whole bunch of other genes, except that it’s not just a single chain of events. There are multiple chains of events, leading to massive disruption of gene expression in autism.”
A 2011 study by Hu’s group revealed that RORA has the potential to be under negative and positive regulation by androgen and estrogen, respectively, suggesting that RORA may also contribute to the male bias of autism spectrum disorder.
Notes about this autism and genetics research
The new study, titled “Genome-wide identification of transcriptional targets of RORA reveals direct regulation of multiple genes associated with autism spectrum disorder,” is available online.
Contact: Lisa Anderson – George Washington University
Source: George Washington University press release
Image Source: The flow diagram image is credited to Valerie W Hu and is adapted from the George Washington University press release.
Original Research: Full open access research for “Genome-wide identification of transcriptional targets of RORA reveals direct regulation of multiple genes associated with autism spectrum disorder” by Tewarit Sarachana and Valerie W Hu in Molecular Autism. Published online May 22 2013 doi:10.1186/2040-2392-4-14