Summary: A new study reveals molecular mechanism that helps insure continuous readjustment of the strenght of synaptic junctions during learning.
Source: LMU.
Learning involves constant restructuring of nerve-cell connections. This requires specialized transport systems to ensure that these specific synapses can structurally and functionally be modified. One such delivery mechanism has now been characterized.
At the cellular level, learning and memory require continuous readjustment of the strength of synaptic junctions – the functional connections between nerve cells. These modifications are achieved by the incorporation into individual synapses of proteins that modulate the efficiency of intercellular signal transmission. The blueprints for the synthesis of these molecules, in the form of messenger RNAs (mRNAs), are delivered to active synapses by a dedicated transport system. An international team led by LMU biochemist Professor Michael Kiebler has now demonstrated that a key factor involved in this transport binds specifically to regions of its mRNA cargo that lack protein-coding information, and that release of the RNAs to enable protein synthesis is dependent on synaptic activity. The new findings appear online in the journal EMBO Reports.
Nerve cells typically consist of a cell body (in which the nucleus resides), a long fiber (the axon) which conducts outgoing nerve impulses, and a set of shorter projections (dendrites) near the cell body that serve as antennas for incoming signals. The instructions for the synthesis of proteins are stored in the DNA in the cell nucleus, are first ‘transcribed’ into mRNAs, and exported from the nucleus to the ribosomes in the cell cytoplasm. Ribosomes are the molecular machines responsible for the translation of mRNA sequences into the amino-acid sequences of the specified newly synthesized protein. In nerve cells, ribosomes are also found close to the synapses of the dendrites, thus enabling proteins that are required for synaptic function to be produced close to their sites of action. This localized delivery system also enables particular synapses to be specifically modified without concomitant alteration of their neighbors.
Kiebler and his group had previously shown that the brain-specific RNA-binding protein Staufen2 (Stau2) plays a central role in the transport of mRNAs from the cell nucleus to synaptic ribosomes. “However, it remained unclear how Stau2 recognizes the mRNAs it conveys to the synapses, and how these are translated into proteins that modify specific synapses in an activity-dependent manner,” Kiebler explains.
In the new study, the researchers show that Stau2 binds specifically to a so-called intron sequence found near the end of a specific subset of mRNAs – in a region that contains no coding information, but serves a regulatory function instead. Introns are regions in the so-called primary transcript transcribed from the nuclear DNA, which are either removed (by a process called alternative splicing) or selectively retained prior to export of the processed mRNA. “Most introns found in mRNAs are located in protein-coding regions and their selective retention permits different versions – so-called isoforms – of a protein to be synthesized from the same precursor RNA. It is however quite unusual to find an intron retained in the untranslated region of an mRNA,” says Tejaswini Sharangdhar, first author of the study. Moreover, Stau2 fails to bind to the mRNA isoform that lacks the intron. Consequently, this RNA isoform is not transported to the dendrites for translation. Thus the intron in the non-coding region serves as one of the regulatory markers which ensure that mRNA is recognized by Stau2 and transported to synapses.
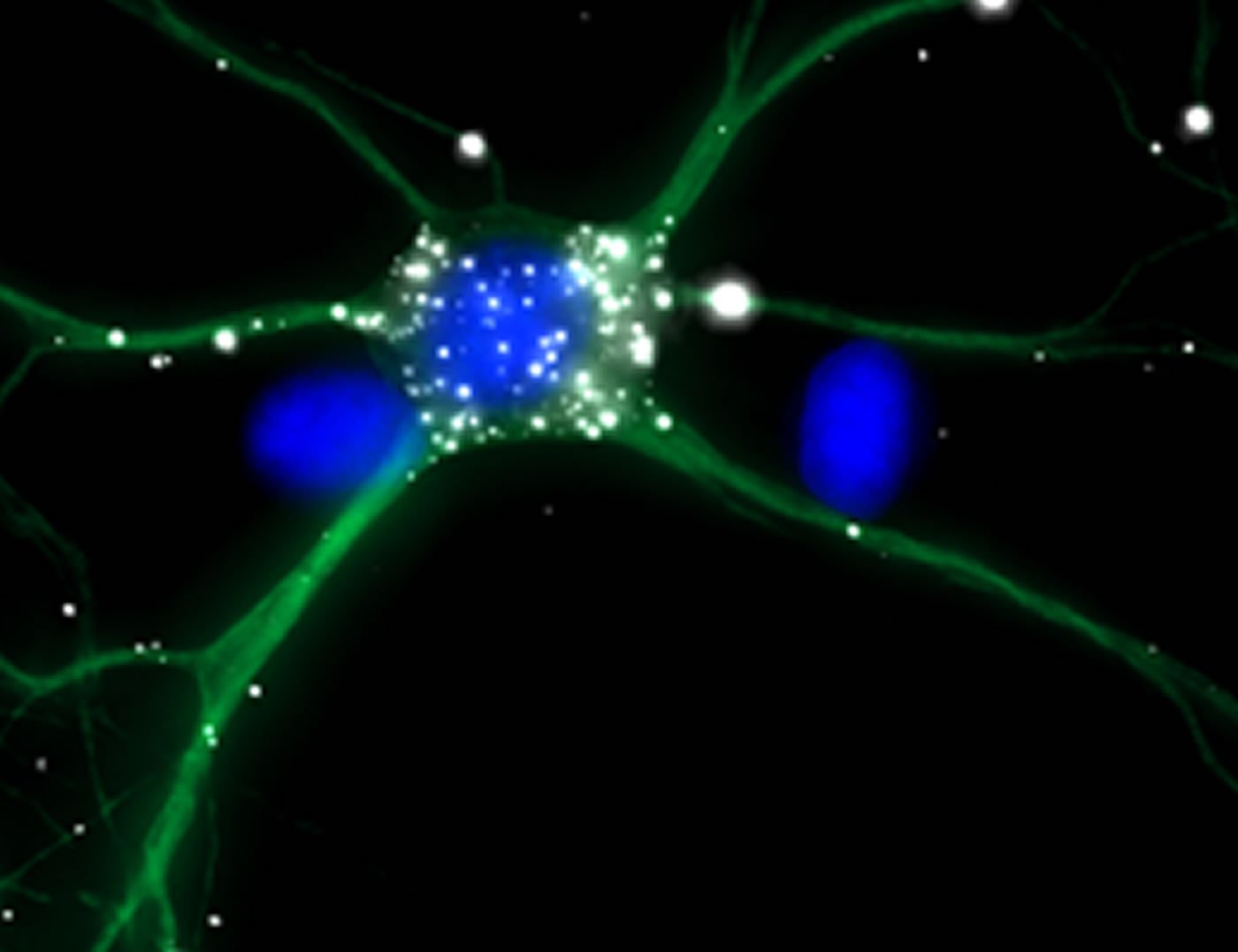

“Our second surprising discovery is that this transport process is dependent on synaptic activity,” says Kiebler: The intron-containing mRNA is transported from one synapse to the next – but will only be released for translation in the vicinity of synapses that are momentarily active. Hence, synaptic activity stimulates the docking of the mRNA to specific synapses, and its protein product marks “learning” synapses. Here too, Stau2 plays an essential role, for in the absence of the protein, none of the mRNA reaches the dendrites.
Source: LMU
Image Source: NeuroscienceNews.com image is credited to T. Sharangdhar and M. Kiebler, LMU.
Original Research: Abstract for “A retained intron in the 3′-UTR of Calm3 mRNA mediates its Staufen2- and activity-dependent localization to neuronal dendrites” by Tejaswini Sharangdhar, Yoichiro Sugimoto, Jacqueline Heraud-Farlow, Sandra M Fernández-Moya, Janina Ehses, Igor Ruiz de los Mozos, Jernej Ule, and Michael A Kiebler in EMBO Reports. Published online August 1 2017 doi:10.15252/embr.201744334
[cbtabs][cbtab title=”MLA”]LMU “Lessons by Post.” NeuroscienceNews. NeuroscienceNews, 7 August 2017.
<https://neurosciencenews.com/learning-pruning-7258/>.[/cbtab][cbtab title=”APA”]LMU (2017, August 7). Lessons by Post. NeuroscienceNew. Retrieved August 7, 2017 from https://neurosciencenews.com/learning-pruning-7258/[/cbtab][cbtab title=”Chicago”]LMU “Lessons by Post.” https://neurosciencenews.com/learning-pruning-7258/ (accessed August 7, 2017).[/cbtab][/cbtabs]
Abstract
A retained intron in the 3′-UTR of Calm3 mRNA mediates its Staufen2- and activity-dependent localization to neuronal dendrites
Dendritic localization and hence local mRNA translation contributes to synaptic plasticity in neurons. Staufen2 (Stau2) is a well-known neuronal double-stranded RNA-binding protein (dsRBP) that has been implicated in dendritic mRNA localization. The specificity of Stau2 binding to its target mRNAs remains elusive. Using individual-nucleotide resolution CLIP (iCLIP), we identified significantly enriched Stau2 binding to the 3′-UTRs of 356 transcripts. In 28 (7.9%) of those, binding occurred to a retained intron in their 3′-UTR. The strongest bound 3′-UTR intron was present in the longest isoform of Calmodulin 3 (Calm3L) mRNA. Calm3L 3′-UTR contains six Stau2 crosslink clusters, four of which are in this retained 3′-UTR intron. The Calm3L mRNA localized to neuronal dendrites, while lack of the 3′-UTR intron impaired its dendritic localization. Importantly, Stau2 mediates this dendritic localization via the 3′-UTR intron, without affecting its stability. Also, NMDA-mediated synaptic activity specifically promoted the dendritic mRNA localization of the Calm3L isoform, while inhibition of synaptic activity reduced it substantially. Together, our results identify the retained intron as a critical element in recruiting Stau2, which then allows for the localization of Calm3L mRNA to distal dendrites.
“A retained intron in the 3′-UTR of Calm3 mRNA mediates its Staufen2- and activity-dependent localization to neuronal dendrites” by Tejaswini Sharangdhar, Yoichiro Sugimoto, Jacqueline Heraud-Farlow, Sandra M Fernández-Moya, Janina Ehses, Igor Ruiz de los Mozos, Jernej Ule, and Michael A Kiebler in EMBO Reports. Published online August 1 2017 doi:10.15252/embr.201744334