Caltech biologists have performed the first large-scale screening in a vertebrate animal for genes that regulate sleep, and have identified a gene that when overactivated causes severe insomnia. Expression of the gene, neuromedin U (Nmu), also seems to serve as nature’s stimulant–fish lacking the gene take longer to wake up in the morning and are less active during the day.
The findings improve our understanding of how sleep is regulated–a process that we know surprisingly little about despite its clear importance. In the long term, the results suggest Nmu as a potential candidate for new therapies to address sleep disorders.
A paper describing the new screening process and its results appears in the February 17, 2016, issue of the journal Neuron. David Prober, assistant professor of biology at Caltech, began the work as a postdoctoral fellow at Harvard University, and has continued the work at Caltech since 2009. The lead authors on the paper are Cindy Chiu (PhD ’14), a former graduate student in Prober’s lab, and Jason Rihel, who collaborated with Prober at Harvard and now has his own lab at University College London.
“Sleep is a mysterious process,” says Prober. “We spend a third of our lives doing it, and every animal with a complex nervous system seems to do it, so it must be important. But we still don’t understand why we do it or how it’s regulated.”
Genetic screens are a powerful method that can help identify the genetic basis of such behaviors. They typically involve mutating the DNA of thousands of animals, raising them, identifying any resulting physical or behavioral differences, and determining which altered gene produced each mutation. This approach works well for simple model organisms, such as fruit flies and worms, because their anatomy is relatively simple and it is easy to raise large numbers of them, but is far more difficult in vertebrates, such as rodents.
Recently, zebrafish have emerged as a valuable vertebrate model system for studying sleep. Compared to a mouse, the small, striped fish are much easier to work with. Many can be raised in a small space (a larva is about 4 millimeters long, about the same size as a fruit fly); they develop quickly, exhibiting complex behaviors, such as hunting, by the time they are five days old; and they are transparent during their embryonic and larval stages, making it simpler for researchers to track what is happening inside their brains. Like humans, zebrafish sleep for consolidated periods of time at night. Furthermore, Prober says, “anatomical and molecular similarities between zebrafish and mammalian brains suggest that the basic neural circuits regulating sleep in zebrafish are likely conserved in mammals.”
Rather than mutating the DNA and testing which functions were lost, the researchers used a gain-of-function approach in the new study. Just after fertilization, when the zebrafish embryos were still single cells, the researchers injected them with a DNA molecule, called a plasmid, carrying a gene that was inserted into the genome of some of the cells in each fish. In particular, they wanted to test genes that are predicted to encode for secreted proteins–those, like neuropeptides, that cells make and then release. Many of the genes that have been identified as being involved in sleep encode neuropeptides.
Using a genetic switch called a heat-shock promoter, which turns on only when the fish are heated to about 37 degrees Celsius, the biologists were able to control when the fish expressed each inserted gene. They kept the switch on long enough for the fish to overexpress each gene, making many copies of the products. Then they used computerized video trackers to monitor the fish for several days to see which genes affected sleep.
Next, the researchers made transgenic zebrafish for each of the genes that had demonstrated strong effects on sleep in the genetic screen. That labor-intensive approach gave them zebrafish in which all cells overexpressed a particular gene in response to heat shock, providing more robust results.
In the end, the most significant change resulted from overexpression of Nmu, a gene that is also found in mammals and is expressed in a part of the brain called the hypothalamus.
“After heat shock, the fish that overexpress Nmu are much more active both during the day and at night,” says Prober. “The fish almost don’t sleep at all the night following the heat shock–so they display a very profound form of insomnia.”
When the researchers mutated the zebrafish so that they did not have Nmu, the larvae were less active during the day. Adult fish without the gene were particularly sluggish first thing in the morning.
Like humans, zebrafish normally start to wake up at the end of the night and then become much more active when the lights turn on. “The fish without Nmu are defective in this anticipation of dawn,” says Prober. “So it seems that this gene is particularly important for the transition from nighttime sleep to daytime wakefulness.”
To explore how Nmu promotes wakefulness, the researchers first investigated the gene’s role in a stress response pathway known as the hypothalamic-pituitary-adrenal (HPA) axis. Researchers had previously shown Nmu to be involved in arousal caused by stressful situations and hypothesized that it was involved in activating the HPA axis. However, Prober and his colleagues found that Nmu suppressed sleep to the same extent in zebrafish mutants lacking a protein called the glucocorticoid receptor, which is necessary for HPA axis signaling, as it did in fish with a functional glucocorticoid receptor, suggesting that the gene does not act through the HPA axis.
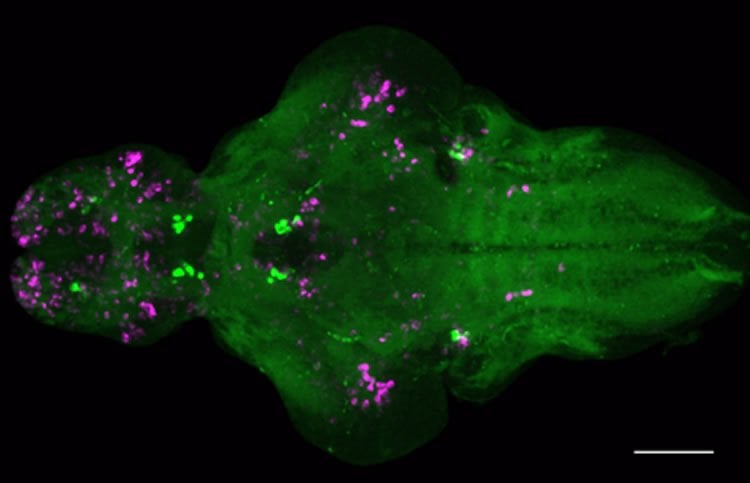
The researchers then went back to the drawing board and asked which neurons in the brain became activated as a result of Nmu overexpression. Using a technique that labels activated neurons, they saw strong activation of a handful of cells that express a gene called corticotrophin-releasing hormone (CRH) in the brainstem.
“That was surprising because CRH is the gene that initiates the HPA axis response, but the cells that do that are in the hypothalamus, a different part of the brain, and they aren’t activated when we overexpress Nmu,” says Prober. “It’s another population of CRH cells in the brainstem that are activated by Nmu overexpression.”
This video shows a larval zebrafish locomotor activity assay that is used to monitor sleep/wake behaviors. Individual five-day old zebrafish, which are about 4 millimeters long, are placed into each well of a 96-well plate, and a computer tracks their behavior for several days.
A low dose of a drug that blocks CRH signaling completely blocked the wake-promoting effect of Nmu overexpression in zebrafish, the researchers found, whereas a higher dose also reduced wakefulness in normal fish.
“So not only is CRH signaling required for the effects of Nmu on behavior, it’s also required for normal levels of activity,” explains Prober.
Several wake-promoting or sleep-promoting genes and neurons have been identified, he notes. However, scientists still do not know which are the relevant ones for causing sleep disorders in humans. “Our study suggests that Nmu could be a good gene to look into.”
Additional Caltech authors on the paper, “A Zebrafish Genetic Screen Identifies Neuromedin U as a Regulator of Sleep/Wake States,” are Daniel A. Lee, Chanpreet Singh, Eric A. Mosser, Shijia Chen, Viveca Sapin, Uyen Pham, Jae Engle, Brett J. Niles, Christin J. Montz, and Sridhara Chakravarthy. Steven Zimmerman and Alexander F. Schier are additional authors from Harvard University. Kourosh Salehi-Ashtiani and Marc Vidal are authors from Harvard Medical School.
Funding: The work was supported by grants from the National Institutes of Health, the European Research Council, University College London, the High-Tech Fund of the Dana Farber Cancer Institute, the Ellison Foundation, the Edward Mallinckrodt, Jr. Foundation, the Rita Allen Foundation, and the Brain and Behavior Research Foundation.
Source: Kimm Fesenmaier – CalTech
Image Source: The image is credited to Cindy N. Chiu and David A. Prober/Caltech.
Video Source: The video is available via the CalTech YouTube page
Original Research: Abstract for “A Zebrafish Genetic Screen Identifies Neuromedin U as a Regulator of Sleep/Wake States” by Cindy N. Chiu, Jason Rihel, Daniel A. Lee, Chanpreet Singh, Eric A. Mosser, Shijia Chen, Viveca Sapin, Uyen Pham, Jae Engle, Brett J. Niles, Christin J. Montz, Sridhara Chakravarthy, Steven Zimmerman, Kourosh Salehi-Ashtiani, Marc Vidal, Alexander F. Schier, and David A. Prober in Neuron. Published online February 17 2016 doi:10.1016/j.neuron.2016.01.007
Abstract
A Zebrafish Genetic Screen Identifies Neuromedin U as a Regulator of Sleep/Wake States
Highlights
•A zebrafish genetic screen identifies genes that regulate sleep and arousal
•Nmu overexpression inhibits sleep whereas nmu mutants are hypoactive
•Nmu-induced phenotypes require Nmu receptor 2 but not the glucocorticoid receptor
•Nmu-induced arousal activates brainstem Crh neurons and requires Crh signaling
Summary
Neuromodulation of arousal states ensures that an animal appropriately responds to its environment and engages in behaviors necessary for survival. However, the molecular and circuit properties underlying neuromodulation of arousal states such as sleep and wakefulness remain unclear. To tackle this challenge in a systematic and unbiased manner, we performed a genetic overexpression screen to identify genes that affect larval zebrafish arousal. We found that the neuropeptide neuromedin U (Nmu) promotes hyperactivity and inhibits sleep in zebrafish larvae, whereas nmu mutant animals are hypoactive. We show that Nmu-induced arousal requires Nmu receptor 2 and signaling via corticotropin releasing hormone (Crh) receptor 1. In contrast to previously proposed models, we find that Nmu does not promote arousal via the hypothalamic-pituitary-adrenal axis, but rather probably acts via brainstem crh-expressing neurons. These results reveal an unexpected functional and anatomical interface between the Nmu system and brainstem arousal systems that represents a novel wake-promoting pathway.
“A Zebrafish Genetic Screen Identifies Neuromedin U as a Regulator of Sleep/Wake States” by Cindy N. Chiu, Jason Rihel, Daniel A. Lee, Chanpreet Singh, Eric A. Mosser, Shijia Chen, Viveca Sapin, Uyen Pham, Jae Engle, Brett J. Niles, Christin J. Montz, Sridhara Chakravarthy, Steven Zimmerman, Kourosh Salehi-Ashtiani, Marc Vidal, Alexander F. Schier, and David A. Prober in Neuron. Published online February 17 2016 doi:10.1016/j.neuron.2016.01.007