Summary: Combining microscopy with artificial intelligence, researchers were able to visualize the complex architecture of interconnected neurons in live C. elegans.
Source: Yale
The formation of a brain is one of nature’s most staggeringly complex accomplishments. The intricate intermingling of neurons and a labyrinth of connections also make it a particularly difficult feat for scientists to study.
Now, Yale researchers and collaborators have devised a strategy that allows them to see this previously impenetrable process unfold in a living animal — the worm Caenorhabditis elegans, they report February 24 in the journal Nature.
“Before, we were able to study single cells, or small groups of cells, in the context of the living C. elegans, and for relatively short periods of time,” said Mark Moyle, an associate research scientist in neuroscience at Yale School of Medicine and first author of the study. “It has been a breathtaking experience to now be able to watch development unfold for hours, across the entire brain of the organism, and visualize this highly orchestrated dance.”
The researchers describe the choreography of a developing brain in this video.
Moyle works in the lab headed by corresponding author Daniel Colón-Ramos, the Dorys McConnell Duberg Professor of Neuroscience and Cell Biology and senior author of the study.
The lab collaborated with computational and microscopy scientists to develop novel network algorithms and imaging technologies that allowed them to study complex webs of interconnected neurons in living C. elegans, a common type of roundworm often used in research. Despite its simplicity, it shares key molecular and genetic characteristics with human biology.
The researchers found that interconnected neurons, densely packed into units called neuropils, are organized to sort signals which dictate many functions and behaviors in the organisms. The study details architectural principles in the neuropil structure that determines how functional brain circuits are developed and assembled.
The authors found that neuronal processes and connections in the worm’s brain are organized into layers, each containing modular components of functional circuits that are linked to distinct behaviors.
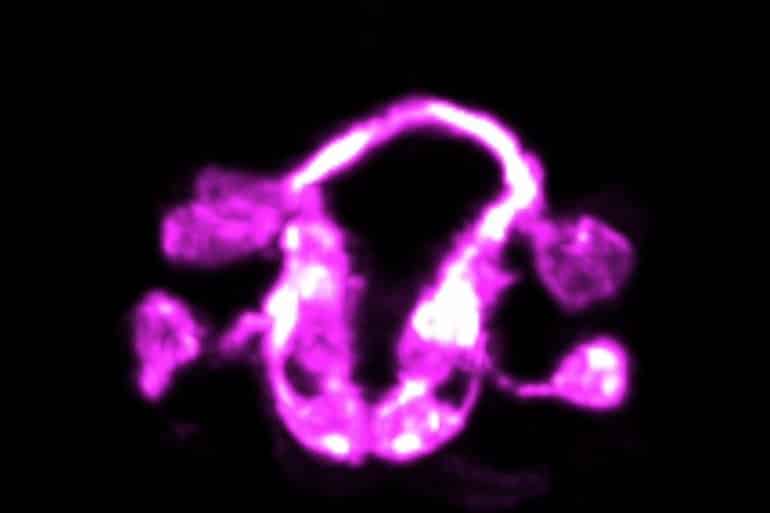
Then, using high-resolution light sheet microscopy, the researchers were able to track single cells over the course of the organism’s development, providing insights into how these cells help choreograph the assembly of the brain.
“When you see the architecture, you realize that all this knowledge that was out there about the animal’s behaviors has a home in the structure of the brain,” Colón-Ramos said.
For instance, researchers can trace reflex behavior in animals to circuits leading to muscles and how these same circuits integrate with still others to regulate the animal’s movement.
He said the brain is organized like a city such as New York, with areas like Wall Street or Broadway organized to carry out the specific functions of finance and entertainment, respectively.
“Suddenly you see how the city fits together and you understand the relationships between the neighborhoods,” Colon-Ramos said.
The work is the result of a decade long collaboration between the labs of Colón-Ramos and Smita Krishnaswamy of Yale; Hari Shroff of the National Institutes of Health; Zhirong Bao of the Sloan Kettering Institute; and William A. Mohler of the University of Connecticut Health Center. The Colón-Ramos, Shroff, Bao, and Mohler labs are part of the consortium WormGUIDES.
About this brain development research news
Source: Yale
Contact: Bess Connolly – Yale
Image: The image is credited to the researchers
Original Research: Closed access.
“Structural and developmental principles of neuropil assembly in C. elegans” by Mark W. Moyle, Kristopher M. Barnes, Manik Kuchroo, Alex Gonopolskiy, Leighton H. Duncan, Titas Sengupta, Lin Shao, Min Guo, Anthony Santella, Ryan Christensen, Abhishek Kumar, Yicong Wu, Kevin R. Moon, Guy Wolf, Smita Krishnaswamy, Zhirong Bao, Hari Shroff, William A. Mohler & Daniel A. Colón-Ramos. Nature
Abstract
Structural and developmental principles of neuropil assembly in C. elegans
Neuropil is a fundamental form of tissue organization within the brain, in which densely packed neurons synaptically interconnect into precise circuit architecture. However, the structural and developmental principles that govern this nanoscale precision remain largely unknown. Here we use an iterative data coarse-graining algorithm termed ‘diffusion condensation’ to identify nested circuit structures within the Caenorhabditis elegans neuropil, which is known as the nerve ring.
We show that the nerve ring neuropil is largely organized into four strata that are composed of related behavioural circuits. The stratified architecture of the neuropil is a geometrical representation of the functional segregation of sensory information and motor outputs, with specific sensory organs and muscle quadrants mapping onto particular neuropil strata.
We identify groups of neurons with unique morphologies that integrate information across strata and that create neural structures that cage the strata within the nerve ring. We use high resolution light-sheet microscopy coupled with lineage-tracing and cell-tracking algorithms to resolve the developmental sequence and reveal principles of cell position, migration and outgrowth that guide stratified neuropil organization.
Our results uncover conserved structural design principles that underlie the architecture and function of the nerve ring neuropil, and reveal a temporal progression of outgrowth—based on pioneer neurons—that guides the hierarchical development of the layered neuropil. Our findings provide a systematic blueprint for using structural and developmental approaches to understand neuropil organization within the brain.