Smart songbird’s reference genome is milestone for ecological research.
A well-known songbird, the great tit, has revealed its genetic code, offering researchers new insight into how species adapt to a changing planet. Their initial findings suggest that epigenetics — what’s on rather than what’s in the gene — may play a key role in the evolution of memory and learning. And that’s not just true for birds. An international research team led by the Netherlands Institute of Ecology (NIOO-KNAW) and Wageningen University will publish these findings in Nature Communications on Monday.
“People in our field have been waiting for this for decades,” explain researchers Kees van Oers and Veronika Laine from the Netherlands Institute of Ecology. The reference genome of their favourite model species, the great tit, is “a powerful toolbox that all ecologists and evolutionary biologists should know about.”
Coming from a single Dutch bird, the genetic code of the assembled reference genome will help to reveal the genetic basis of phenotypic evolution. This is essential for understanding how wild species adapt to our changing planet.
In addition to looking at the genome, the research team have also determined the so-called transcriptome and methylome. The latter belongs to the field of epigenetics: the study of what you can inherit not in but ‘on’ your genes. Specific DNA sequences in the genome can be ‘methylated’: methyl groups are added to them, modifying how the genes function.
The research team sequenced the complete genomes of a further 29 great tit individuals from different parts of Europe. This enabled them to identify regions in the great tit’s genome that have been under selection during recent evolution of the bird. These regions appeared to be overrepresented for genes related to learning and cognition.
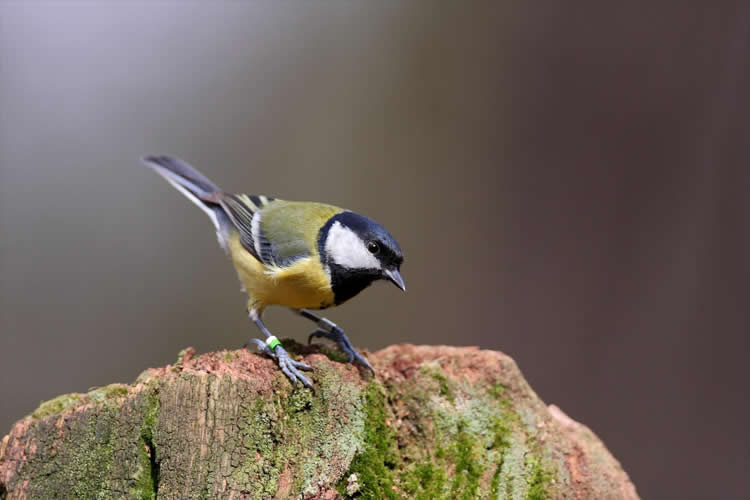

“The great tit has evolved to be smart,” says Van Oers. “Very smart.” It’s not your average bird, as it belongs to the top 3% smartest birds when it comes to learning new behaviour. That makes it a perfect candidate for research into the evolution of learning, memory and cognitive processes.
What that research has revealed are so-called conserved patterns of methylation in those same regions, present not only in birds but also in humans and other mammals. It’s evidence of a correlation between epigenetic processes such as methylation and the rate of molecular evolution: “the more methylation, the more evolution.”
And so the great tit has once more proved that its role as a model species in a variety of biological research fields for over 60 years is by no means coincidental.”
The international team that completed the genome sequence of the great tit consisted of scientists from four European institutes (NIOO-KNAW, Wageningen University, University of Sheffield and University of Oxford), two from the USA (University of Illinois and Washington University School of Medicine) and a consortium of other great tit researchers from around the world.
Funding: The research was supported by the Rural Development Administration Republic of Korea, Biotechnology and Biological Sciences Research Council, New World Order, Royal Society.
Source: Froukje Rienks – Netherlands Institute of Ecology
Image Credit: The image is credited to Netherlands Institute of Ecology (NIOO-KNAW)
Original Research: Full open access research for “Evolutionary signals of selection on cognition from the great tit genome and methylome” by Veronika N. Laine, Toni I. Gossmann, Kyle M. Schachtschneider, Colin J. Garroway, Ole Madsen, Koen J. F. Verhoeven, Victor de Jager, Hendrik-Jan Megens, Wesley C. Warren, Patrick Minx, Richard P. M. A. Crooijmans, Pádraic Corcoran, The Great Tit HapMap Consortium, Ben C. Sheldon, Jon Slate, Kai Zeng, Kees van Oers, Marcel E. Visser and Martien A. M. Groenen in Nature Communications. Published online January 25 2016 doi:10.1038/ncomms10474
Abstract
Evolutionary signals of selection on cognition from the great tit genome and methylome
For over 50 years, the great tit (Parus major) has been a model species for research in evolutionary, ecological and behavioural research; in particular, learning and cognition have been intensively studied. Here, to provide further insight into the molecular mechanisms behind these important traits, we de novo assemble a great tit reference genome and whole-genome re-sequence another 29 individuals from across Europe. We show an overrepresentation of genes related to neuronal functions, learning and cognition in regions under positive selection, as well as increased CpG methylation in these regions. In addition, great tit neuronal non-CpG methylation patterns are very similar to those observed in mammals, suggesting a universal role in neuronal epigenetic regulation which can affect learning-, memory- and experience-induced plasticity. The high-quality great tit genome assembly will play an instrumental role in furthering the integration of ecological, evolutionary, behavioural and genomic approaches in this model species.
“Evolutionary signals of selection on cognition from the great tit genome and methylome” by Veronika N. Laine, Toni I. Gossmann, Kyle M. Schachtschneider, Colin J. Garroway, Ole Madsen, Koen J. F. Verhoeven, Victor de Jager, Hendrik-Jan Megens, Wesley C. Warren, Patrick Minx, Richard P. M. A. Crooijmans, Pádraic Corcoran, The Great Tit HapMap Consortium, Ben C. Sheldon, Jon Slate, Kai Zeng, Kees van Oers, Marcel E. Visser and Martien A. M. Groenen in Nature Communications. Published online January 25 2016 doi:10.1038/ncomms10474