University of Utah chemists look for precursors to disease mutations.
University of Utah chemists devised a new way to detect chemical damage to DNA that sometimes leads to genetic mutations responsible for many diseases, including various cancers and neurological disorders.
“We are one step closer to understanding the underlying chemistry that leads to genetic diseases,” says Cynthia Burrows, distinguished professor and chair of chemistry at the university and senior author of a new study published Nov. 6 in the online journal Nature Communications. “We have a way of marking and copying DNA damage sites so that we can preserve the information of where and what the damage was.”
Jan Riedl, a University of Utah postdoctoral fellow and the study’s first author, says 99 percent of DNA lesions – damage to the chemical bases known as A, C, G and T that help form the DNA double helix – are repaired naturally. The rest can lead to genetic mutations, which are errors in the sequence of bases and can cause disease. The new method can “identify and detect the position of lesions that lead to diseases,” he says.
Burrows says: “We are trying to look for the chemical changes in the base that can lead the cell to make a mistake, a mutation. One of the powerful things about our method is we can read more than a single damaged site [and up to dozens] on the same strand of DNA.”
The chemists say their new method will let researchers study chemical details of DNA lesions or damage. Such lesions, if not repaired naturally, accumulate over time and can lead to mutations responsible for many age-related diseases, including colon, breast, liver, lung and melanoma skin cancers; clogged arteries; and neurological ailments such as Huntington’s disease and Lou Gehrig’s disease.
“A method capable of identifying the chemical identity and location in which lesions appear is crucial for determining the molecular etiology [cause] of these diseases,” Burrows and colleague write in their study.
The research was funded by the National Institutes of Health. Burrows and Riedl conducted the study with University of Utah chemists Aaron Fleming, a research assistant professor, and Yun Ding, a postdoctoral researcher.
Method detects DNA damage that can lead to disease
DNA is a molecule, shaped like a double-stranded helix that contains the genetic instructions that living organisms use to develop, function and reproduce. Each strand is made of numerous copies of four chemical bases – A, C, G and T – linked by a backbone of sugar and phosphate. When two strands are connected in a double helix, only two kinds of base pairs can form C-G (or G-C) and A-T (or T-A).
The new method for finding DNA lesions combines other, existing techniques.
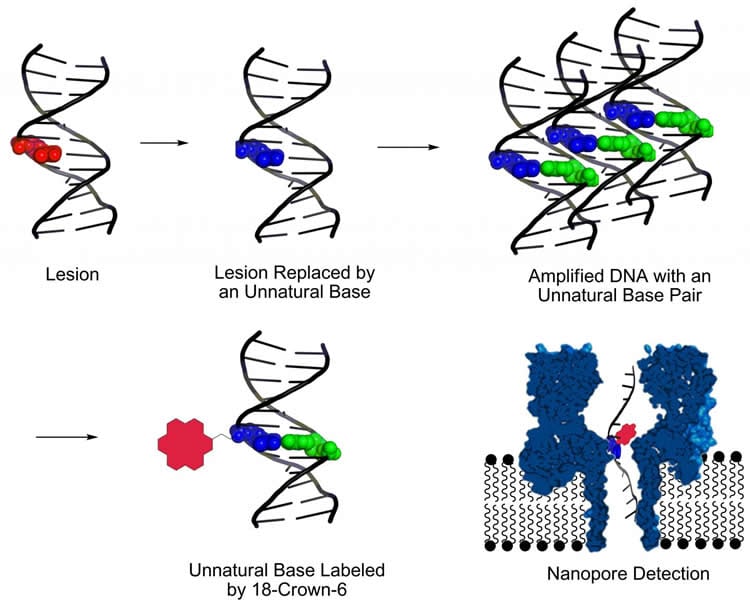
First, the researchers find the damage and cut it out of the DNA the same way a cell does naturally, using what is called “base excision repair,” the discovery of which won a Nobel Prize in Chemistry this year for Tomas Lindahl, a scientist in England.
Second, an “unnatural base pair” is inserted at the snipped-out DNA damage site to label it. Instead of natural base pairs C-G and A-T, the Utah chemists used a so-called third or unnatural base pair invented by chemists at the Scripps Research Institute in California. Burrows says her study demonstrates the first practical use of that invention.
Third, the DNA with the damage site labeled by an unnatural third base pair is then amplified or copied millions of times using a well-known existing method called PCR, or polymerase chain reaction. Burrows says the new study’s key innovation was to use base excision repair to snip out the damage and then to insert the unnatural base pair at the damage site, making it possible to make millions of copies of the DNA – a process that normally would be prevented by the damage.
Fourth, another chemical label, named 18-crown-6 ether, is affixed to the unnatural base pair on all the DNA strands, which are then read or sequenced using a kind of nanopore sequencing developed a few years ago by Burrows and Utah chemist Henry White. Such sequencing involves determining the order and location of bases on a DNA strand – including damage sites labeled by unnatural bases – by passing the strand through a molecule-size pore or nanopore.
People are born with their genome or genetic blueprint of 3 billion base pairs, “and then stuff happens,” Burrows says. “There’s damage from oxidative stress due to inflammation and infection, too much metabolism, or environmental chemicals.”
The new method seeks “molecular details that define how our genome responds to what we eat and the air we breathe, and ends up being healthy or not,” she says.
DNA lesions happen more than 10,000 times a day in a single human cell. A lesion can be a missing base, a base that has changed chemically or a break in the DNA backbone. That many lesions may seem like a lot, but with 3 billion base pairs in the genome of a single cell, the damage only affects about one of every 300,000 base pairs.
Details of the study
The chemists tested their method on a gene named KRAS that, when mutated, can cause lung or breast cancer.
Burrows says most DNA sequencing methods reveal mutations because the methods read the bases A, C, G and T. When sequencing reveals one of those bases out of place, that is a mutation. “However, what you don’t know is what chemistry – what modification – caused that mutation,” she says,
But scientists can’t find a single strand of DNA in a solution in a reasonable time. They need to make millions to billions of copies of the DNA so they can sequence it and locate the gap where the damage was. But the damage itself is “a train wreck” that either prevents making copies of the DNA or makes copies with errors, Burrows says.
So once the damaged base has been excised to create a gap, the chemists insert the third or unnatural base pair at then damage site as a way to label it while at the same time allowing millions of copies of the labeled DNA to be made.
Burrows says the key innovation in the new method is using base excision repair and unnatural base pairs “to copy the DNA and retain the information about damage that was in the original molecule.”
The chemists next use PCR amplification to make millions of copies of the DNA by heating it until strands in the double helix separate. The strands are put in a solution with lots of A, C, G and T nucleotides – the bases attached to pieces of DNA backbone. A polymerase enzyme is attached to the end of each strand of DNA, and then moves along the strand grabbing T, G, C and A nucleotides to make a second strand. Each DNA strand quickly becomes two. The number of strands reaches the millions in hours.
“We need that many because, if we just have one molecule, we can’t find it,” Burrows says. “It’s like saying, ‘There’s one fish in that pond; try to catch it.’ But if there are a bunch of identical fish, you can catch any one of them and still have dinner.”
Once the chemists have millions of DNA strands with the damage labeled by an unnatural base pair, they then use nanopore sequencing to locate the damage. Burrows says to imagine the DNA strand is a long piece of yarn with a number of knots tied in it to represent labeled DNA lesions. Grab one end of the yarn with one hand, and pull it through a couple of fingers on the other hand. The fingers represent the nanopore that detects the knots or damage as they pass between the fingers.
To do nanopore sequencing, the DNA’s damage sites, which are labeled with unnatural bases, are labeled again with a chemical named 18-crown-6 ether that basically makes the “knots” on the DNA “yarn” bigger and easier to find as the DNA strand moves through a pore-shaped protein bathed in electrically charged fluid. Changes in electric current allow chemists detect the bases A, C, G and T and the labeled unnatural bases.
Finding the labeled unnatural bases identifies the DNA damage site to within 10 base pairs. Burrows says the nanopore sequencing method needs improvement or replacement by next-generation sequencing to actually pinpoint the damage sites.
Funding: The study was funded by the National Institutes of Health.
Source: Lee J. Siegel – University of Utah
Image Credit: The image is credited to Aaron Fleming, University of Utah
Original Research: Full open access research for “Identification of DNA lesions using a third base pair for amplification and nanopore sequencing” by Jan Riedl, Yun Ding, Aaron M. Fleming and Cynthia J. Burrows in Nature Communications. Published online November 62015 doi:10.1038/ncomms9807
Abstract
Identification of DNA lesions using a third base pair for amplification and nanopore sequencing
Damage to the genome is implicated in the progression of cancer and stress-induced diseases. DNA lesions exist in low levels, and cannot be amplified by standard PCR because they are frequently strong blocks to polymerases. Here, we describe a method for PCR amplification of lesion-containing DNA in which the site and identity could be marked, copied and sequenced. Critical for this method is installation of either the dNaM or d5SICS nucleotides at the lesion site after processing via the base excision repair process. These marker nucleotides constitute an unnatural base pair, allowing large quantities of marked DNA to be made by PCR amplification. Sanger sequencing confirms the potential for this method to locate lesions by marking, amplifying and sequencing a lesion in the KRAS gene. Detection using the α-hemolysin nanopore is also developed to analyse the markers in individual DNA strands with the potential to identify multiple lesions per strand.
“Identification of DNA lesions using a third base pair for amplification and nanopore sequencing” by Jan Riedl, Yun Ding, Aaron M. Fleming and Cynthia J. Burrows in Nature Communications. Published online November 62015 doi:10.1038/ncomms9807