Summary: Researchers present a new approach to creating genetic knock out mice to help investigate circadian rhythm.
Source: RIKEN.
A new study from the laboratory of Hiroki Ueda at the RIKEN Quantitative Biology Center investigates circadian timekeeping with a novel approach to creating genetic knock-out rescue mice. Published in Molecular Cell, the study shows how this technique was used to quickly create numerous mouse lines, each with different mutations in a circadian regulator called CRY1. Studying each mutation and the effects on behavior showed that specific changes to the protein affected the duration of the circadian period.
Circadian rhythms are physiological changes at the cellular levels in almost all plants and animals that follow a 24-hour cycle. They are guided by internal biological clocks, and are affected by many internal genetics factors, which in turn can influence behavior.
Next-generation mammalian genetics is a specialty of Ueda and his team. Recently, they succeeded in efficiently creating gene knockout (KO) mice without backcrossing or breeding. Until now however, the inverse procedures called knock-in (KI) and KO-rescue have been laborious, time-consuming, and costly. Ueda and his team have now developed a method for efficiency producing KI and KO-rescue that involves using three-inhibitor (3i) treatment of mouse embryonic stem cells. With this procedure, they can begin with KO embryonic stem cells, insert rescue genes, and then analyze the behavior of the mouse when it matures–all within a single generation.
The group used this new KO-rescue technique to better understand cryptochromes–proteins associated with physiological functions that are controlled by environmental light. While cryptochromes are related to circadian rhythms in both plants and animals, their exact physiological roles appear quite different. In mammals, cryptochrome gene CRY1 is essential for a normal circadian rhythm, but its exact role in regulating the length of circadian periods remains unclear. Guided by mass spectrometry-based identification of certain sites at CRY1, and comprehensive analysis of all CRY1 mutants at each site, Ueda and his team identified more than ten areas in the gene that affect the period of the circadian clock in cultured cells.
Ueda and his team selected 17 CRY1 mutations, and added each mutant CRY1 into different mouse embryonic stem cells that lacked all cryptochrome genes. When the mice grew up, they then analyzed their activity patterns. “Mice without these genes lack a circadian rhythm,” explains first author Koji Ode. “By adding the Cry1 gene back in these mice, we were able to rescue the circadian rhythm. Most interestingly, the lengths of the restored circadian periods depended on nature of the mutations.”
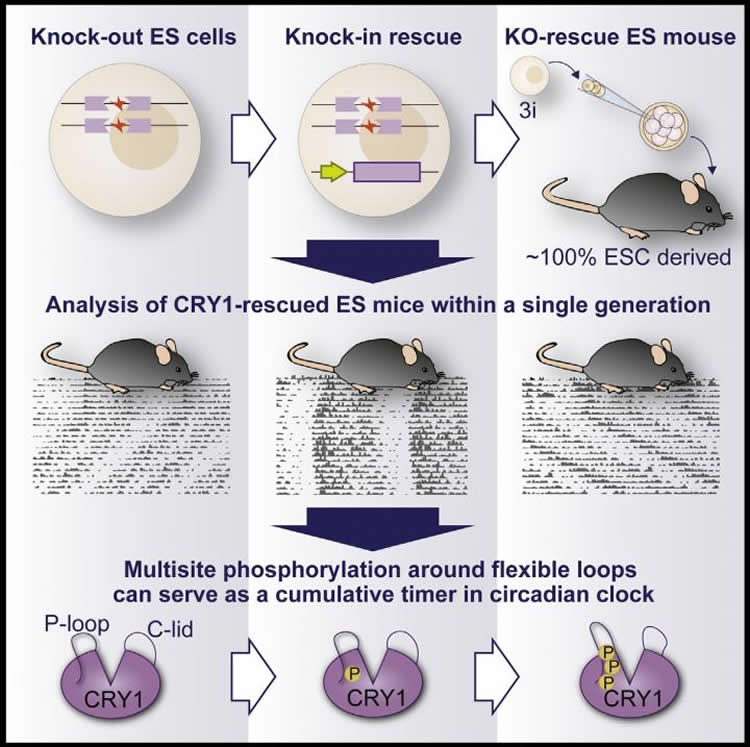
The most effective versions of the Cry1 genes to be knocked-in had a mutation near the p-loop, which codes for a pocket-like area on the protein that can be modified through a process called phosphorylation–the attachment of phosphate groups. The KO-rescue mice with mutations that limited the amount of phosphorylation around the CRY1 p-loop had longer than normal circadian periods.
“This p-loop is critical for CRY1 to function as a circadian clock” says Ueda. “We think that the accumulation of phosphorylation at this site serves as a time-keeping mechanism and thus the quality of the p-loop is a fundamental element that regulates circadian rhythm. Our next challenge is to actually observe changes in phosphorylation level on the p-loop over the course of a day.”
Source: Adam Phillips – RIKEN
Image Source: NeuroscienceNews.com image is credited to RIKEN.
Original Research: Abstract for “Knockout-Rescue Embryonic Stem Cell-Derived Mouse Reveals Circadian-Period Control by Quality and Quantity of CRY1” by Koji L. Ode, Hideki Ukai, Etsuo A. Susaki, Ryohei Narumi, Katsuhiko Matsumoto, Junko Hara, Naoshi Koide, Takaya Abe, Masato T. Kanemaki, Hiroshi Kiyonari, and Hiroki R. Ueda in Molecular Cell. Published online December 22 2016 doi:10.1016/j.molcel.2016.11.022
[cbtabs][cbtab title=”MLA”]RIKEN “Controlling the Body Clock.” NeuroscienceNews. NeuroscienceNews, 22 December 2016.
<https://neurosciencenews.com/cry1-circadian-rhythm-5803/>.[/cbtab][cbtab title=”APA”]RIKEN (2016, December 22). Controlling the Body Clock. NeuroscienceNew. Retrieved December 22, 2016 from https://neurosciencenews.com/cry1-circadian-rhythm-5803/[/cbtab][cbtab title=”Chicago”]RIKEN “Controlling the Body Clock.” https://neurosciencenews.com/cry1-circadian-rhythm-5803/ (accessed December 22, 2016).[/cbtab][/cbtabs]
Abstract
Knockout-Rescue Embryonic Stem Cell-Derived Mouse Reveals Circadian-Period Control by Quality and Quantity of CRY1
Highlights
•A KO-rescue ES mouse method was developed to produce 20 different KO-rescue strains
•Multisite phosphorylation of CRY1 can serve as a cumulative timer
•CRY1-PER2 interaction confers a robust circadian rhythmicity in mice
•Flexible loops of CRY1 determine circadian period in mice without a turnover change
Summary
To conduct comprehensive characterization of molecular properties in organisms, we established an efficient method to produce knockout (KO)-rescue mice within a single generation. We applied this method to produce 20 strains of almost completely embryonic stem cell (ESC)-derived mice (“ES mice”) rescued with wild-type and mutant Cry1 gene under a Cry1−/−:Cry2−/− background. A series of both phosphorylation-mimetic and non-phosphorylation-mimetic CRY1 mutants revealed that multisite phosphorylation of CRY1 can serve as a cumulative timer in the mammalian circadian clock. KO-rescue ES mice also revealed that CRY1-PER2 interaction confers a robust circadian rhythmicity in mice. Surprisingly, in contrast to theoretical predictions from canonical transcription/translation feedback loops, the residues surrounding the flexible P loop and C-lid domains of CRY1 determine circadian period without changing the degradation rate of CRY1. These results suggest that CRY1 determines circadian period through both its degradation-dependent and -independent pathways.
“Knockout-Rescue Embryonic Stem Cell-Derived Mouse Reveals Circadian-Period Control by Quality and Quantity of CRY1” by Koji L. Ode, Hideki Ukai, Etsuo A. Susaki, Ryohei Narumi, Katsuhiko Matsumoto, Junko Hara, Naoshi Koide, Takaya Abe, Masato T. Kanemaki, Hiroshi Kiyonari, and Hiroki R. Ueda in Molecular Cell. Published online December 22 2016 doi:10.1016/j.molcel.2016.11.022