Findings point to potential therapeutic targets.
Using advanced DNA sequencing methods, researchers have identified a new gene that is associated with sporadic amyotrophic lateral sclerosis (ALS), or Lou Gehrig’s disease. ALS is a devastating neurodegenerative disorder that results in the loss of all voluntary movement and is fatal in the majority of cases. The next-generation genetic sequencing of the exomes (protein-coding portions) of 2,874 ALS patients and 6,405 controls represents the largest number of ALS patients to have been sequenced in a single study to date.
Though much is known about the genetic underpinnings of familial ALS, only a handful of genes have been definitively linked to sporadic ALS, which accounts for about 90 percent of all ALS cases. The newly associated gene, called TBK1, plays a key role at the intersection of two essential cellular pathways: inflammation (a reaction to injury or infection) and autophagy (a cellular process involved in the removal of damaged cellular components). The study, conducted by an international ALS consortium that includes scientists and clinicians from Columbia University Medical Center (CUMC), Biogen Idec, and HudsonAlpha Institute for Biotechnology, was published today in the online edition of Science.
“The identification of TBK1 is exciting for understanding ALS pathogenesis, especially since the inflammatory and autophagy pathways have been previously implicated in the disease,” said Lucie Bruijn, PhD, Chief Scientist for The ALS Association. “The fact that TBK1 accounts for one percent of ALS adds significantly to our growing understanding of the genetic underpinnings of the disease. This study, which combines the efforts of over two dozen laboratories in six countries, also highlights the global and collaborative nature of ALS research today.
“This study shows us that large-scale genetic studies not only can work very well in ALS, but that they can help pinpoint key biological pathways relevant to ALS that then become the focus of targeted drug development efforts,” said study co-leader David B. Goldstein, PhD, professor of genetics and development and director of the new Institute for Genomic Medicine at CUMC. “ALS is an incredibly diverse disease, caused by dozens of different genetic mutations, which we’re only beginning to discover. The more of these mutations we identify, the better we can decipher–and influence–the pathways that lead to disease.” The other co-leaders of the study are Richard M. Myers, PhD, president and scientific director of HudsonAlpha, and Tim Harris, PhD, DSc, Senior Vice President, Technology and Translational Sciences, Biogen Idec.
“These findings demonstrate the power of exome sequencing in the search for rare variants that predispose individuals to disease and in identifying potential points of intervention. We are following up by looking at the function of this pathway so that one day this research may benefit the patients living with ALS,” said Dr. Harris. “The speed with which we were able to identify this pathway and begin our next phase of research shows the potential of novel, focused collaborations with the best academic scientists to advance our understanding of the molecular pathology of disease. This synergy is vital for both industry and the academic community, especially in the context of precision medicine and whole-genome sequencing.”
“Industry and academia often do things together, but this is a perfect example of a large, complex project that required many parts, with equal contributions from Biogen Idec. Dr. Tim Harris, our collaborator there, and his team, as well as David Goldstein and his team, now at Columbia University, as well as our teams here at HudsonAlpha, said Dr. Myers. “I love this research model because it doesn’t happen very frequently, and it really shows how industry, nonprofits, and academic laboratories can all work together for the betterment of humankind. The combination of those groups with a large number of the clinical collaborators who have been seeing patients with this disease for many years and providing clinical information, recruiting patients, as well as collecting DNA samples for us to do this study, were all critical to get this done.”
Searching through the enormous database generated in the ALS study, Dr. Goldstein and his colleagues found several genes that appear to contribute to ALS, most notably TBK1 (TANK-Binding Kinase 1), which had not been detected in previous, smaller-scale studies. TBK1 mutations appeared in about 1 percent of the ALS patients–a large proportion in the context of a complex disease with multiple genetic components, according to Dr. Goldstein. The study also found that a gene called OPTN, previously thought to play a minor role in ALS, may actually be a major player in the disease.
“Remarkably, the TBK1 protein and optineurin, which is encoded by the OPTN gene, interact physically and functionally. Both proteins are required for the normal function of inflammatory and autophagy pathways, and now we have shown that mutations in either gene are associated with ALS,” said Dr. Goldstein. “Thus there seems to be no question that aberrations in the pathways that require TBK1 and OPTN are important in some ALS patients.”
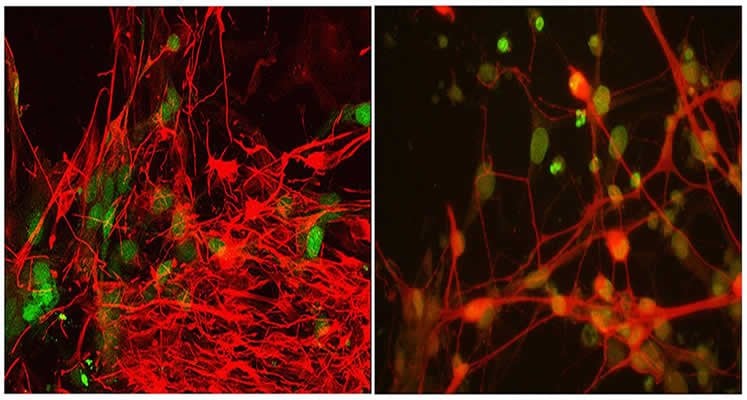
The researchers are currently using patient-derived induced pluripotent embryonic stem cells (iPS cells) and mouse models with mutations in TBK1 or OPTN to study ALS disease mechanisms and to screen for drug candidates. Several compounds that affect TBK1 signaling have already been developed for use in cancer, where the gene is thought to play a role in tumor-cell survival.
“This is a great example of the potential of precision medicine,” said Tom Maniatis, PhD, the Isidore S. Edelman Professor, chair of biochemistry and molecular biophysics, and coauthor on the paper. Dr. Maniatis is also a member of the Zuckerman Mind Brain Behavior Institute and director of Columbia’s university-wide precision medicine initiative. “It now seems clear that future ALS treatments will not be equally effective for all patients because of the disease’s genetic diversity. Ultimately, as candidate therapies become available, we hope to be able to use the genetic data from each ALS patient to direct that person to the most appropriate clinical trials and, ultimately, use the data to prescribe treatment.”
The study was funded by Biogen Idec, Wellcome Trust and Medical Research Council Grants, MND Association, the ALS Association, National Institutes of Health (R37 NS083524, 5RO1NS050557, 5R01 NS067206, 5R01NS065847, 1R01NS073873, 5R01NS079836, R01NS065847, and R01NS073873), the Angel Fund, Project ALS/P2ALS, the ALS Therapy Alliance, the Pierre L. de Bourghknecht ALS Research Foundation, a Francesco Caleffi donation, the Medical Research Council, the Heaton-Ellis Trust and AriSLA cofinanced with support of ”5 x 1000”–Healthcare research of the Italian Ministry of Health, the National Institute for Health Research Dementia Biomedical Research Unit at South London, Maudsley NHS Foundation Trust, King’s College London, the Motor Neurone Disease Research Institute of Australia, United Kingdom Medical Research Council under the aegis of JPND, the National Health and Medical Research Council of Australia, Canadian Institutes of Health Research, Muscle Dystrophy Association, Instituto de Salud Carlos III – ISCIII, FUNDELA (Spanish foundation for the development of ALS research), the Mireia Barneda project, the Netherlands ALS Foundation, and the European Community’s Health Seventh Framework Programme.
J. Wade Harper is a consultant for Biogen Idec and Millennium: the Takeda Oncology Company. Robert Brown is a consultant for Biogen Idec and a cofounder of AviTx. Frank Baas is a founder of Regenesance. The other authors report no conflict of interest.
Contact: Karin Eskenazi – Columbia Univeresity Medical Center
Source: Columbia Univeresity Medical Center press release
Image Source: The image is credited to Laboratory of Tom Maniatis and is adapted from the Columbia Univeresity Medical Center press release
Original Research: Abstract for “Exome sequencing in amyotrophic lateral sclerosis identifies risk genes and pathways” by Elizabeth T. Cirulli, Brittany N. Lasseigne, Slavé Petrovski, Peter C. Sapp, Patrick A. Dion, Claire S. Leblond, Julien Couthouis, Yi-Fan Lu, Quanli Wang, Brian J. Krueger, Zhong Ren, Jonathan Keebler, Yujun Han, Shawn E. Levy, Braden E. Boone, Jack R. Wimbish, Lindsay L. Waite, Angela L. Jones, John P. Carulli, Aaron G. Day-Williams, John F. Staropoli, Winnie W. Xin, Alessandra Chesi, Alya R. Raphael, Diane McKenna-Yasek, Janet Cady, J. M. B. Vianney de Jong, Kevin P. Kenna, Bradley N. Smith, Simon Topp, Jack Miller, Athina Gkazi, FALS Sequencing Consortium, Ammar Al-Chalabi, Leonard H. van den Berg, Jan Veldink, Vincenzo Silani, Nicola Ticozzi, Christopher E. Shaw, Robert H. Baloh, Stanley Appel, Ericka Simpson, Clotilde Lagier-Tourenne, Stefan M. Pulst, Summer Gibson, John Q. Trojanowski, Lauren Elman, Leo McCluskey, Murray Grossman, Neil A. Shneider, Wendy K. Chung, John M. Ravits, Jonathan D. Glass, Katherine B. Sims, Vivianna M. Van Deerlin, Tom Maniatis, Sebastian D. Hayes, Alban Ordureau, Sharan Swarup, John Landers, Frank Baas, Andrew S. Allen, Richard S. Bedlack, J. Wade Harper, Aaron D. Gitler, Guy A. Rouleau, Robert Brown, Matthew B. Harms, Gregory M. Cooper, Tim Harris, Richard M. Myers, and David B. Goldstein in Science. Published online February 19 2015 doi:10.1126/science.aaa3650