Benjamin Blencowe, a professor in the University of Toronto’s Donnelly Centre and Banbury Chair in Medical Research, and his team have uncovered how a small change in a protein called PTBP1 can spur the creation of neurons – cells that make the brain – that could have fuelled the evolution of mammalian brains to become the largest and most complex among vertebrates.
The study is published in the August 20 issue of Science.
Brain size and complexity vary enormously across vertebrates, but it is not clear how these differences came about. Humans and frogs, for example, have been evolving separately for 350 million years and have very different brain abilities. Yet scientists have shown that they use a remarkably similar repertoire of genes to build organs in the body.
So how is it that a similar number of genes, that are also switched on or off in similar ways in diverse vertebrate species, generate a vast range of organ size and complexity?
The key lays in the process that Blencowe’s group studies, known as alternative splicing (AS), whereby gene products are assembled into proteins, which are the building blocks of life. During AS, gene fragments – called exons – are shuffled to make different protein shapes. It’s like LEGO, where some fragments can be missing from the final protein shape.
AS enables cells to make more than one protein from a single gene, so that the total number of different proteins in a cell greatly surpasses the number of available genes. A cell’s ability to regulate protein diversity at any given time reflects its ability to take on different roles in the body. Blencowe’s previous work showed that AS prevalence increases with vertebrate complexity. So although the genes that make bodies of vertebrates might be similar, the proteins they give rise to are far more diverse in animals such as mammals, than in birds and frogs.
And nowhere is AS more widespread than in the brain.
“We wanted to see if AS could drive morphological differences in the brains of different vertebrate species,” says Serge Gueroussov, a graduate student in Blencowe’s lab who is the lead author of the study. Gueroussov previously helped identify PTBP1 as a protein that takes on another form in mammals, in addition to the one common to all vertebrates. The second form of mammalian PTBP1 is shorter because a small fragment is omitted during AS and does not make it into the final protein shape.
Could this newly acquired, mammalian version of PTBP1 give clues to how our brains evolved?
PTBP1 is both a target and major regulator of AS. PTBP1’s job in a cell is to stop it from becoming a neuron by holding off AS of hundreds of other gene products.
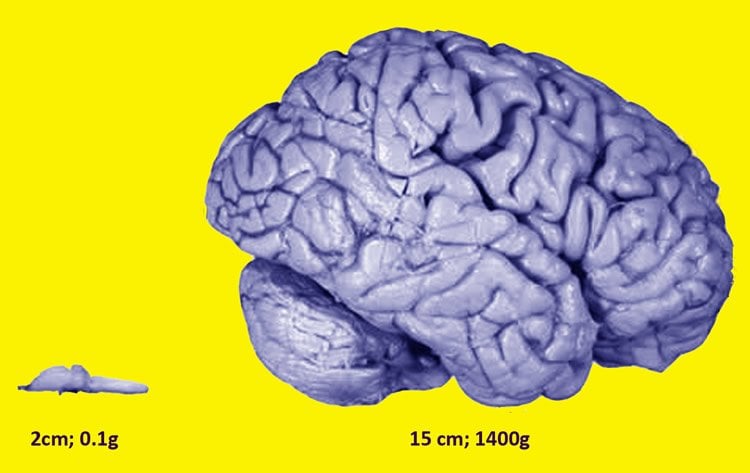
Gueroussov showed that in mammalian cells, the presence of the second, shorter version of PTBP1 unleashes a cascade of AS events, tipping the scales of protein balance so that a cell becomes a neuron.
What’s more, when Gueroussov engineered chicken cells to make the shorter, mammalian-like, PTBP1, this triggered AS events that are found in mammals.
“One interesting implication of our work is that this particular switch between the two versions of PTBP1 could have affected the timing of when neurons are made in the embryo in a way that creates differences in morphological complexity and brain size,” says Blencowe, who is also a professor in the Department of Molecular Genetics.
As scientists continue to sift through countless molecular events occurring in our cells, they’ll keep finding clues as to how our bodies and minds came to be.
“This is the tip of an iceberg in terms of the full repertoire of AS changes that likely have contributed major roles in driving evolutionary differences,” says Blencowe.
Source: Jovana Drinjakovic – University of Toronto
Image Credit: The image is credited to the researchers/University of Toronto
Original Research: Abstract for “An alternative splicing event amplifies evolutionary differences between vertebrates” by Serge Gueroussov, Thomas Gonatopoulos-Pournatzis, Manuel Irimia, Bushra Raj, Zhen-Yuan Lin, Anne-Claude Gingras, and Benjamin J. Blencowe in Science. Published online August 21 2015 doi:10.1126/science.aaa8381
Abstract
An alternative splicing event amplifies evolutionary differences between vertebrates
Alternative splicing (AS) generates extensive transcriptomic and proteomic complexity. However, the functions of species- and lineage-specific splice variants are largely unknown. Here we show that mammalian-specific skipping of polypyrimidine tract–binding protein 1 (PTBP1) exon 9 alters the splicing regulatory activities of PTBP1 and affects the inclusion levels of numerous exons. During neurogenesis, skipping of exon 9 reduces PTBP1 repressive activity so as to facilitate activation of a brain-specific AS program. Engineered skipping of the orthologous exon in chicken cells induces a large number of mammalian-like AS changes in PTBP1 target exons. These results thus reveal that a single exon-skipping event in an RNA binding regulator directs numerous AS changes between species. Our results further suggest that these changes contributed to evolutionary differences in the formation of vertebrate nervous systems.
“An alternative splicing event amplifies evolutionary differences between vertebrates” by Serge Gueroussov, Thomas Gonatopoulos-Pournatzis, Manuel Irimia, Bushra Raj, Zhen-Yuan Lin, Anne-Claude Gingras, and Benjamin J. Blencowe in Science. Published online August 21 2015 doi:10.1126/science.aaa8381