DNA mutations can cause cancer but in some cases, more mutations may mean a better prognosis for patients.
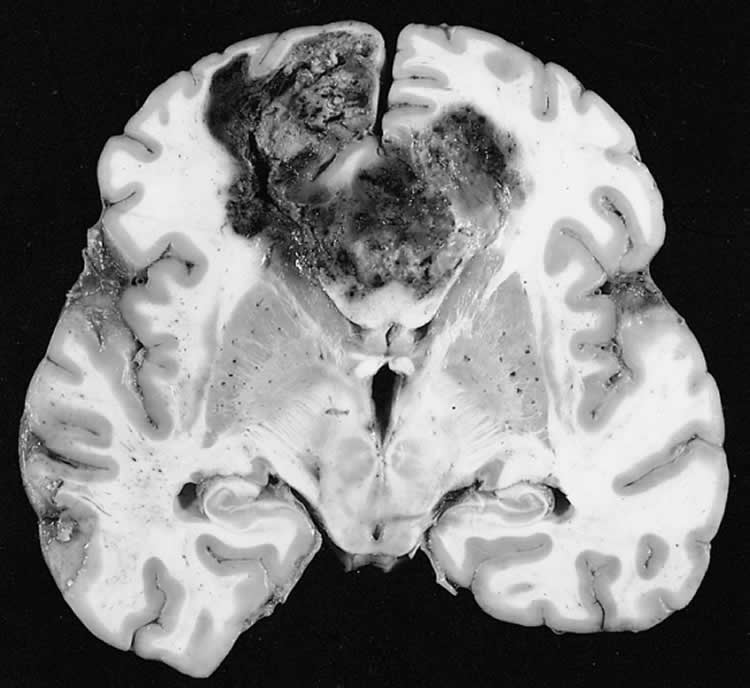
A Yale-led comprehensive genomic analysis of more than 700 brain tumors has revealed one such subtype of the most malignant brain tumor, called glioblastoma, or GBM. This subtype possesses thousands of tumor-specific DNA errors or mutations instead of dozens observed in most glioblastoma cases. It is also associated with longer survival.
The findings, reported in the journal, Neuro-Oncology, suggest it may be possible to develop personalized treatments for more aggressive forms of brain cancer, including immunotherapy for these hyper- or ultra-mutated tumors, said Murat Günel, professor and chair of neurosurgery, who leads the Brain Tumor Research Program at Yale and Smilow Cancer Hospital at Yale-New Haven Hospital.
“We have been able to translate various complementary cutting-edge genomic technologies, which were once solely research tools, to our clinical programs to analyze individual cancers,” said Günel, who is also a professor of genetics and a researcher for the Yale Cancer Center. “We can now gain comprehensive understanding of the molecular make-up of a cancer to pinpoint specific vulnerabilities and leverage these weak spots for precision treatments in our Recurrent Brain Tumor Treatment Program.”
While as many as 10,000 mutations were found in the newly described subset of glioblastomas, a more typical tumor contains less than 100. This counterintuitive pattern has also been observed in gynecological and colon cancers: An extraordinary number of mutations means a better chance of survival.
One theory holds that cells with greater number of mutations are able to trigger an aggressive immune system response against cancer cells, while cells with fewer mutations might escape detection, Gunel said.
Although the number of GBMs in this newly identified group is small, the use of standard chemotherapy in some cases has been shown to inadvertently result in a hyper-mutated tumor. Indeed, the drug temozolomide, used as the first line of chemotherapy in GBM, has been shown to sometimes increase mutations.
“But perhaps the naïve immune system is not strong enough to eliminate the cancer cells in these brain tumors,” Gunel noted.
However, if a new generation of immunotherapy drugs called checkpoint inhibitors were used in these hyper-mutated tumors, perhaps more cancer cells might be targeted for destruction, he said. Clinical trials currently underway might be improved by considering the molecular genetic make-up of the individual tumor, he concluded.
Zeynep Erson-Omay and Ahmet Okay Çağlayan from Yale co-first authored the research paper.
Funding: The Gregory Kiez and Mehmet Kutman Foundation funded the work.
Article written by Bill Hathaway
Source: Yale News
Research Article: Abstract for “Somatic POLE mutations cause an ultramutated giant cell high-grade glioma subtype with better prognosis” by E. Zeynep Erson-Omay†, Ahmet Okay Çağlayan†, Nikolaus Schultz, Nils Weinhold, S. Bülent Omay, Koray Özduman, Yavuz Köksal, Jie Li, Akdes Serin Harmancı, Victoria Clark, Geneive Carrión-Grant, Jacob Baranoski, Caner Çağlar, Tanyeri Barak, Süleyman Coşkun, Burçin Baran, Doğan Köse, Jia Sun, Mehmet Bakırcıoğlu, Jennifer Moliterno Günel, M. Necmettin Pamir, Ketu Mishra-Gorur, Kaya Bilguvar, Katsuhito Yasuno, Alexander Vortmeyer, Anita J. Huttner, Chris Sander and Murat Günel in Neuro- Oncology first published online March 3, 2015 doi: 10.1093/neuonc/nov02
Related Resource: Yale Program in Brain Tumor Research
Abstract
Background
Malignant high-grade gliomas (HGGs), including the most aggressive form, glioblastoma multiforme, show significant clinical and genomic heterogeneity. Despite recent advances, the overall survival of HGGs and their response to treatment remain poor. In order to gain further insight into disease pathophysiology by correlating genomic landscape with clinical behavior, thereby identifying distinct HGG molecular subgroups associated with improved prognosis, we performed a comprehensive genomic analysis.
Methods
We analyzed and compared 720 exome-sequenced gliomas (136 from Yale, 584 from The Cancer Genome Atlas) based on their genomic, histological, and clinical features.
Results
We identified a subgroup of HGGs (6 total, 4 adults and 2 children) that harbored a statistically significantly increased number of somatic mutations (mean = 9257.3 vs 76.2, P = .002). All of these “ultramutated” tumors harbored somatic mutations in the exonuclease domain of the polymerase epsilon gene (POLE), displaying a distinctive genetic profile, characterized by genomic stability and increased C-to-A transversions. Histologically, they all harbored multinucleated giant or bizarre cells, some with predominant infiltrating immune cells. One adult and both pediatric patients carried homozygous germline mutations in the mutS homolog 6 (MSH6) gene. In adults, POLE mutations were observed in patients younger than 40 years and were associated with a longer progression-free survival.
Conclusions
We identified a genomically, histologically, and clinically distinct subgroup of HGGs that harbored somatic POLE mutations and carried an improved prognosis. Identification of distinctive molecular and pathological HGG phenotypes has implications not only for improved classification but also for potential targeted treatments.