Computer modeling identifies first-ever molecule to inhibit a transient cellular event that drives glioblastoma, and the molecule shrinks glioblastoma in mice.
Patients with glioblastoma, a type of malignant brain tumor, usually survive fewer than 15 months following diagnosis. Since there are no effective treatments for the deadly disease, University of California, San Diego researchers developed a new computational strategy to search for molecules that could be developed into glioblastoma drugs. In mouse models of human glioblastoma, one molecule they found shrank the average tumor size by half. The study is published October 30 by Oncotarget.
The newly discovered molecule works against glioblastoma by wedging itself in the temporary interface between two proteins whose binding is essential for the tumor’s survival and growth. This study is the first to demonstrate successful inhibition of this type of protein, known as a transcription factor.
“Most drugs target stable pockets within proteins, so when we started out, people thought it would be impossible to inhibit the transient interface between two transcription factors,” said first author Igor Tsigelny, PhD, research scientist at UC San Diego Moores Cancer Center, as well as the San Diego Supercomputer Center and Department of Neurosciences at UC San Diego. “But we addressed this challenge and created a new strategy for drug design — one that we expect many other researchers will immediately begin implementing in the development of drugs that target similar proteins, for the treatment of a variety of diseases.”
Transcription factors control which genes are turned “on” or “off” at any given time. For most people, transcription factors labor ceaselessly in a highly orchestrated system. In glioblastoma, one misfiring transcription factor called OLIG2 keeps cell growth and survival genes “on” when they shouldn’t be, leading to quick-growing tumors.
In order to work, transcription factors must buddy up, with two binding to each other and to DNA at same time. If any of these associations are disrupted, the transcription factor is inhibited.
In this study, Tsigelny and team aimed to disrupt the OLIG2 buddy system as a potential treatment for glioblastoma. Based on the known structure of related transcription factors, study co-author Valentina Kouznetsova, PhD, associate project scientist at UC San Diego, developed a computational strategy to search databases of 3D molecular structures for those small molecules that might engage the hotspot between two OLIG2 transcription factors. The team used the Molecular Operation Environment (MOE) program produced by the Chemical Computing Group in Montreal, Canada and high-performance workstations at the San Diego Supercomputer Center to run the search.
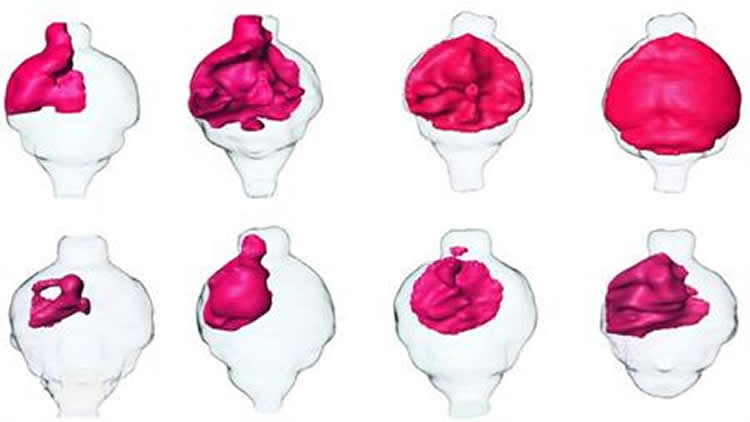
With this approach, the researchers identified a few molecules that would likely fit the OLIG2 interaction. They then tested the molecules for their ability to kill glioblastoma tumors in the Moores Cancer Center lab of the study’s senior author, Santosh Kesari, MD, PhD. The most effective of these candidate drug molecules, called SKOG102, shrank human glioblastoma tumors grown in mouse models by an average of 50 percent.
“While the initial pre-clinical findings are promising,” Kesari cautioned, “it will be several years before a potential glioblastoma therapy can be tested in humans. SKOG102 must first undergo detailed pharmacodynamic, biophysical and mechanistic studies in order to better understand its efficacy and possible toxicity.”
To this end, SKOG102 has been licensed to Curtana Pharmaceuticals, which is currently developing the inhibitor for clinical applications. Kesari is a co-founder, has an equity interest in and is chair of the scientific advisory board for Curtana Pharmaceuticals. Co-authors Rajesh Mukthavaram, PhD, and Wolfgang Wrasidlo, PhD, also own stock in Curtana Pharmaceuticals.
Additional co-authors of this study include Ying Chao, Ivan Babic, Sandra Pastorino, Pengfei Jiang, Miriam Scadeng, Sandeep C. Pingle, Milan T. Makale, UC San Diego; Elmar Nurmemmedov, The Scripps Research Institute; David Calligaris, Nathalie Agar, Harvard Medical School and Brigham and Women’s Hospital.
Funding: This research was funded, in part, by the National Institutes of Health (grant 3P30CA023100-25S8), Voices Against Brain Cancer Foundation, Christopher and Bronwen Gleeson Family Trust and American Brain Tumor Association Drug Discovery Grant.
Source: Heather Buschman – UCSD
Image Source: The image is credited to the researchers/UCSD
Original Research: Abstract for “Multiple spatially related pharmacophores define small molecule inhibitors of OLIG2 in glioblastoma” by Igor F. Tsigelny, Rajesh Mukthavaram, Valentina L. Kouznetsova, Ying Chao, Ivan Babic, Elmar Nurmemmedov, Sandra Pastorino, Pengfei Jiang, David Calligaris, Nathalie Agar, Miriam Scadeng, Sandeep C. Pingle, Wolfgang Wrasidlo, Milan T. Makale and Santosh Kesari in Oncotarget. Published online August 26 2015 doi:10.18632/oncotarget.5633
Abstract
Multiple spatially related pharmacophores define small molecule inhibitors of OLIG2 in glioblastoma
Transcription factors (TFs) are a major class of protein signaling molecules that play key cellular roles in cancers such as the highly lethal brain cancer—glioblastoma (GBM). However, the development of specific TF inhibitors has proved difficult owing to expansive protein-protein interfaces and the absence of hydrophobic pockets. We uniquely defined the dimerization surface as an expansive parental pharmacophore comprised of several regional daughter pharmacophores. We targeted the OLIG2 TF which is essential for GBM survival and growth, we hypothesized that small molecules able to fit each subpharmacophore would inhibit OLIG2 activation. The most active compound was OLIG2 selective, it entered the brain, and it exhibited potent anti-GBM activity in cell-based assays and in pre-clinical mouse orthotopic models. These data suggest that (1) our multiple pharmacophore approach warrants further investigation, and (2) our most potent compounds merit detailed pharmacodynamic, biophysical, and mechanistic characterization for potential preclinical development as GBM therapeutics.
“Multiple spatially related pharmacophores define small molecule inhibitors of OLIG2 in glioblastoma” by Igor F. Tsigelny, Rajesh Mukthavaram, Valentina L. Kouznetsova, Ying Chao, Ivan Babic, Elmar Nurmemmedov, Sandra Pastorino, Pengfei Jiang, David Calligaris, Nathalie Agar, Miriam Scadeng, Sandeep C. Pingle, Wolfgang Wrasidlo, Milan T. Makale and Santosh Kesari in Oncotarget. Published online August 26 2015 doi:10.18632/oncotarget.5633






