Summary: Researchers have developed a scalable method to identify different subtypes of neurons in the brain.
Source: UCSD.
A team of researchers has developed the first scalable method to identify different subtypes of neurons in the human brain. The research lays the groundwork for “mapping” the gene activity in the human brain and could help provide a better understanding of brain functions and disorders, including Alzheimer’s, Parkinson’s, schizophrenia and depression.
By isolating and analyzing the nuclei of individual human brain cells, researchers identified 16 neuronal subtypes in the cerebral cortex — the brain’s outer layer of neural tissue responsible for cognitive functions including memory, attention and decision making. The team, led by researchers at the University of California San Diego, The Scripps Research Institute (TSRI) and Illumina, published their findings in the June 24 online issue of the journal Science.
“We’re providing a unified framework to look at and compare individual neurons, which can help us find out how many unique types of neurons exist,” said Kun Zhang, bioengineering professor at the University of California, San Diego and a corresponding author of the study.
Researchers can use these different neuronal subtypes to build what Zhang calls a “reference map” of the human brain — a foundation to understand the differences between a healthy brain and a diseased brain.
“In the future, patients with brain disorders or abnormalities could be diagnosed and treated based on how they differ from the reference map. This is analogous to what’s being done with the reference human genome map,” Zhang said.
The new study reflects a growing understanding that individual brain cells are unique: they express different types of genes and perform different functions. To better understand this diversity, researchers analyzed more than 3,200 single human neurons in six Brodmann areas, which are regions of the cerebral cortex classified by their functions and arrangements of neurons.
Through an interdisciplinary collaborative effort, the team developed a new method to isolate and sequence individual cell nuclei. TSRI researchers led by neuroscience professor Jerold Chun obtained the samples from a post mortem brain and focused on isolating the neuronal nuclei. Zhang’s lab worked with Fluidigm, a manufacturer of microfluidic chips for single-cell studies, to develop a protocol to identify and quantify RNA molecules in individual neuronal nuclei. Scientists at San Diego-based Illumina sequenced the resulting RNA libraries. Researchers led by biochemistry professor Wei Wang at UC San Diego developed algorithms to cluster and identify 16 neuronal subtypes from the sequenced datasets.
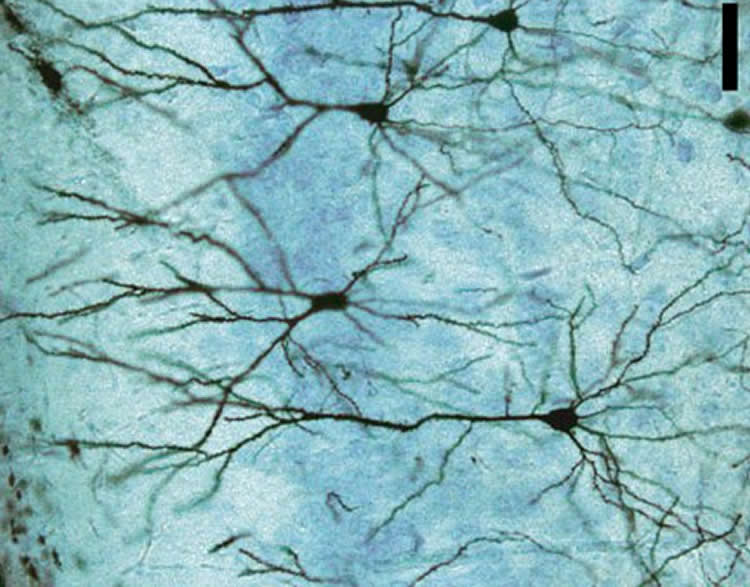
Researchers deciphered what types of genes were “turned on” within each nucleus and revealed that various combinations of the 16 subtypes tended to cluster in cortical layers and Brodmann areas, helping explain why these regions look and function differently.
Neurons exhibited many differences in their transcriptomic profiles — the patterns of genes that are being actively expressed by these cells — revealing single neurons with shared, as well as unique, characteristics that likely lead to difference in cellular function.
“We’re finding new ways to understand the basic building blocks of the brain,” said Blue Lake, a postdoctoral researcher in Zhang’s lab and a co-first author of the study. “Our study opens the door to look at global gene expression patterns and how that defines cell types within a normal tissue, which can also be used to see what’s abnormal in terms of disease or disorders.”
In future studies, researchers aim to analyze neurons in other Brodmann areas of the brain and investigate what subtypes exist in other brain regions. They also plan to study neurons from multiple post mortem human brains (this study only involved one) to investigate neuronal diversity among individuals.
Funding: The interdisciplinary research is part of a larger effort under the NIH Single Cell Analysis Program-Transcriptome (SCAP-T) Project — funded by the NIH Common Fund Program — to develop new single cell analysis technologies.
Source: Liezel Labios – UCSD
Image Source: This NeuroscienceNews.com image is in the public domain.
Original Research: Abstract for “Neuronal subtypes and diversity revealed by single-nucleus RNA sequencing of the human brain” by Blue B. Lake, Rizi Ai, Gwendolyn E. Kaeser, Neeraj S. Salathia, Yun C. Yung, Rui Liu, Andre Wildberg, Derek Gao, Ho-Lim Fung, Song Chen, Raakhee Vijayaraghavan, Julian Wong, Allison Chen, Xiaoyan Sheng, Fiona Kaper, Richard Shen, Mostafa Ronaghi, Jian-Bing Fan, Wei Wang, Jerold Chun, and Kun Zhang in Science. Published online June 23 2016 doi:10.1126/science.aaf1204
[cbtabs][cbtab title=”MLA”]UCSD. “Human Brain Houses Diverse Populations of Neurons.” NeuroscienceNews. NeuroscienceNews, 23 June 2016.
<https://neurosciencenews.com/neuron-diversity-neuroscience-4550/>.[/cbtab][cbtab title=”APA”]UCSD. (2016, June 23). Human Brain Houses Diverse Populations of Neurons. NeuroscienceNews. Retrieved June 23, 2016 from https://neurosciencenews.com/neuron-diversity-neuroscience-4550/[/cbtab][cbtab title=”Chicago”]UCSD. “Human Brain Houses Diverse Populations of Neurons.” https://neurosciencenews.com/neuron-diversity-neuroscience-4550/ (accessed June 23, 2016).[/cbtab][/cbtabs]
Abstract
Neuronal subtypes and diversity revealed by single-nucleus RNA sequencing of the human brain
The human brain has enormously complex cellular diversity and connectivities fundamental to our neural functions, yet difficulties in interrogating individual neurons has impeded understanding of the underlying transcriptional landscape. We developed a scalable approach to sequence and quantify RNA molecules in isolated neuronal nuclei from a postmortem brain, generating 3227 sets of single-neuron data from six distinct regions of the cerebral cortex. Using an iterative clustering and classification approach, we identified 16 neuronal subtypes that were further annotated on the basis of known markers and cortical cytoarchitecture. These data demonstrate a robust and scalable method for identifying and categorizing single nuclear transcriptomes, revealing shared genes sufficient to distinguish previously unknown and orthologous neuronal subtypes as well as regional identity and transcriptomic heterogeneity within the human brain.
“Neuronal subtypes and diversity revealed by single-nucleus RNA sequencing of the human brain” by Blue B. Lake, Rizi Ai, Gwendolyn E. Kaeser, Neeraj S. Salathia, Yun C. Yung, Rui Liu, Andre Wildberg, Derek Gao, Ho-Lim Fung, Song Chen, Raakhee Vijayaraghavan, Julian Wong, Allison Chen, Xiaoyan Sheng, Fiona Kaper, Richard Shen, Mostafa Ronaghi, Jian-Bing Fan, Wei Wang, Jerold Chun, and Kun Zhang in Science. Published online June 23 2016 doi:10.1126/science.aaf1204






