The process that allows our brains to learn and generate new memories also leads to degeneration as we age, according to a new study by researchers at MIT.
The finding, reported in a paper published today in the journal Cell, could ultimately help researchers develop new approaches to preventing cognitive decline in disorders such as Alzheimer’s disease.
Each time we learn something new, our brain cells break their DNA, creating damage that the neurons must immediately repair, according to Li-Huei Tsai, the Picower Professor of Neuroscience and director of the Picower Institute for Learning and Memory at MIT.
This process is essential to learning and memory. “Cells physiologically break their DNA to allow certain important genes to be expressed,” Tsai says. “In the case of neurons, they need to break their DNA to enable the expression of early response genes, which ultimately pave the way for the transcriptional program that supports learning and memory, and many other behaviors.”
Slower DNA repair
However, as we age, our cells’ ability to repair this DNA damage weakens, leading to degeneration, Tsai says. “When we are young, our brains create DNA breaks as we learn new things, but our cells are absolutely on top of this and can quickly repair the damage to maintain the functionality of the system,” Tsai says. “But during aging, and particularly with some genetic conditions, the efficiency of the DNA repair system is compromised, leading to the accumulation of damage, and in our view this could be very detrimental.”
In previous research into Alzheimer’s disease in mice, the researchers found that even in the presymptomatic phase of the disorder, neurons in the hippocampal region of the brain contain a large number of DNA lesions, known as double strand breaks.
To determine how and why these double strand breaks are generated, and what genes are affected by them, the researchers began to investigate what would happen if they created such damage in neurons. They applied a toxic agent to the neurons known to induce double strand breaks, and then harvested the RNA from the cells for sequencing.
They discovered that of the 700 genes that showed changes as a result of this damage, the vast majority had reduced expression levels, as expected. Surprisingly though, 12 genes — known to be those that respond rapidly to neuronal stimulation, such as a new sensory experience — showed increased expression levels following the double strand breaks.
To determine whether these breaks occur naturally during neuronal stimulation, the researchers then treated the neurons with a substance that causes synapses to strengthen in a similar way to exposure to a new experience.
“Sure enough, we found that the treatment very rapidly increased the expression of those early response genes, but it also caused DNA double strand breaks,” Tsai says.
The good with the bad
In further studies the researchers were able to confirm that an enzyme known as topoisomerase IIβ is responsible for the DNA breaks in response to stimulation, according to the paper’s lead author Ram Madabhushi, a postdoc in Tsai’s laboratory.
“When we knocked down this enzyme, we found that both double strand break formation and the expression of early response genes was reduced,” Madabhushi says.
Finally, the researchers attempted to determine why the genes need such a drastic mechanism to allow them to be expressed. Using computational analysis, they studied the DNA sequences near these genes and discovered that they were enriched with a motif, or sequence pattern, for binding to a protein called CTCF. This “architectural” protein is known to create loops or bends in DNA.
In the early-response genes, the bends created by this protein act as a barrier that prevents different elements of DNA from interacting with each other — a crucial step in the genes’ expression.
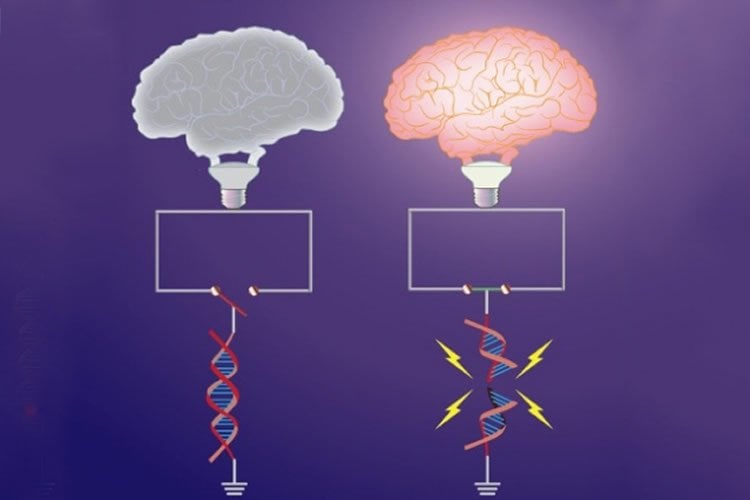
The double strand breaks created by the cells allow them to collapse this barrier, and enable the early response genes to be expressed, Tsai says.
“Surprisingly then, even though conventional wisdom dictates that DNA lesions are very bad — as this ‘damage’ can be mutagenic and sometimes lead to cancer — it turns out that these breaks are part of the physiological function of the cell,” Tsai says.
Previous research has shown that the expression of genes involved in learning and memory is reduced as people age. So the researchers now plan to carry out further studies to determine how the DNA repair system is altered with age, and how this compromises the ability of cells to cope with the continued production and repair of double strand breaks.
They also plan to investigate whether certain chemicals could enhance this DNA repair capacity.
The paper represents an important conceptual advance in our understanding of gene regulation, according to Bruce Yankner, a professor of genetics and neurology at Harvard Medical School who was not involved in the research.
“The work elegantly links DNA strand break formation by the enzyme topoisomerase IIβ to the temporal control of transcription, providing the most compelling evidence to date that this is a core transcriptional control mechanism,” he says. “I anticipate that this advance will have broad implications ranging from the basic biology of transcription to pathological mechanisms involved in diseases such as Alzheimer’s disease.”
Source: Helen Knight – MIT
Image Credit: The image is credited to the researchers
Original Research: Abstract for “Activity-Induced DNA Breaks Govern the Expression of Neuronal Early-Response Genes” by Ram Madabhushi, Fan Gao, Andreas R. Pfenning, Ling Pan, Satoko Yamakawa, Jinsoo Seo, Richard Rueda, Trongha X. Phan, Hidekuni Yamakawa, Ping-Chieh Pao, Ryan T. Stott, Elizabeta Gjoneska, Alexi Nott, Sukhee Cho, Manolis Kellis, and Li-Huei Tsai in Cell. Published online June 4 2015 doi:10.1016/j.cell.2015.05.032
Abstract
Activity-Induced DNA Breaks Govern the Expression of Neuronal Early-Response Genes
Highlights
•Neuronal activity causes the formation of DNA double strand breaks (DSBs)
•Activity-induced DSBs form within the promoters of a subset of early-response genes
•Topoisomerase IIβ is necessary for activity-induced DSB formation
•Activity-induced DSBs facilitate the expression of early-response genes in neurons
Summary
Neuronal activity causes the rapid expression of immediate early genes that are crucial for experience-driven changes to synapses, learning, and memory. Here, using both molecular and genome-wide next-generation sequencing methods, we report that neuronal activity stimulation triggers the formation of DNA double strand breaks (DSBs) in the promoters of a subset of early-response genes, including Fos, Npas4, and Egr1. Generation of targeted DNA DSBs within Fos and Npas4 promoters is sufficient to induce their expression even in the absence of an external stimulus. Activity-dependent DSB formation is likely mediated by the type II topoisomerase, Topoisomerase IIβ (Topo IIβ), and knockdown of Topo IIβ attenuates both DSB formation and early-response gene expression following neuronal stimulation. Our results suggest that DSB formation is a physiological event that rapidly resolves topological constraints to early-response gene expression in neurons.
“Activity-Induced DNA Breaks Govern the Expression of Neuronal Early-Response Genes” by Ram Madabhushi, Fan Gao, Andreas R. Pfenning, Ling Pan, Satoko Yamakawa, Jinsoo Seo, Richard Rueda, Trongha X. Phan, Hidekuni Yamakawa, Ping-Chieh Pao, Ryan T. Stott, Elizabeta Gjoneska, Alexi Nott, Sukhee Cho, Manolis Kellis, and Li-Huei Tsai in Cell. Published online June 4 2015 doi:10.1016/j.cell.2015.05.032