Summary: Researchers have developed a new approach to imaging the functional activity of fetal brains, allowing a better look at how functional brain networks develop.
Source: NIH/NIBIB.
Method makes imaging of moving subjects possible.
NIBIB-funded researchers at the University of Washington have pioneered an approach to image functional activity in the brains of individual fetuses, allowing a better look at how functional networks within the brain develop. The work addresses a common problem of functional MRI; if the subject moves during the scanning, the images get distorted.
By creating a way to correct for motion, the team was able to make a four-dimensional reconstruction of brain activity in moving subjects, opening the door for studies on subjects that don’t stay still for long, like fetuses and small children. The new strategy enables investigations into both normal brain development and the effects of a mother’s diet or environment on the functional development of the fetal brain.
The new work focused on the default mode network—a collection of regions that are active when the brain is at rest, when someone is daydreaming or letting their mind wander, not concentrating on a specific task. Fetal brains are in default mode for much of the time, but it is not very well known how this network develops.
“This is one of the first papers to take individual fetuses and look at the naturally developing default mode network in the human fetal brain,” says Vinay Pai, Ph.D., director of the Division of Health Informatics Technologies at NIBIB. Since movement of the baby or the mother is no longer an issue, “It allows you to look at more natural developmental stages.”
In the new study, published online in August 2016 in the journal Human Brain Mapping, researchers aimed to create a four-dimensional movie of fetal default mode network activity. To do this, they used functional MRI—a technique that detects brain activity based on blood flow; active regions have higher levels of blood flowing to them. Besides blurring the image, the problem with movement—either from the fetus moving or the mother’s breathing—is that it shifts the focus, making it hard to locate where any signals of activity are coming from. While typically just one measurement is collected for each position at each time point, the team collected multiple measurements, each providing slightly different perspectives. Using the multiple measurements, they were able to reposition the images to create an estimate of what the activation over a period of a few minutes would look like.
The team first tested their method on adults by telling them to purposely move their head in the scanner. After showing they could successfully quantify brain activity in moving subjects, they then scanned eight fetuses between 32 and 37 weeks of pregnancy, as infants born prematurely at that age have been shown to have active default mode networks. The resulting images were compiled to create a four-dimensional view of each of the brains over a five-minute time window.
“It really hasn’t been explored when these activity networks—these collections of brain areas that start to work together in the brain—emerge and what types of cells and tissues they emerge in,” says Colin Studholme, Ph.D., a professor with joint appointments in pediatrics and bioengineering at the University of Washington and senior author of the paper. “What this is leading to is not just collecting data from individual babies but also understanding and building a four-dimensional map of brain activity and how it should emerge in a normal baby.”
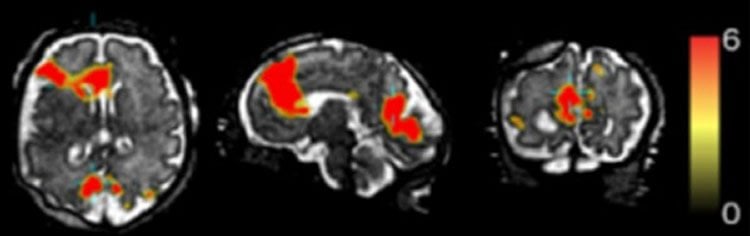
NeuroscienceNews.com image is credited to S. Seshamani, et al.
The technique can also be used to compare differences in brain development in premature and full-term babies; the effects of alcohol, drug use, or stress during pregnancy; or if there are any prenatal differences in babies that go on to develop neurodevelopmental disorders like autism. And it is not restricted to imaging the brain; Studholme also plans to study the placenta and how its development influences the brain.
But the immediate application—for squirmy babies—is promising in itself. “Techniques that minimize the effect of motion resolve a lot of problems,” says Pai. “In fetal imaging, you don’t have too many options.”
Funding: The team received support for this study from NIBIB (EB017133), the National Institue of Neurological Disorders and Stroke (NS055064), and the National Center for Advancing Translational Sciences (UL1TR000423).
Source: Teal Burrell – NIH/NIBIB
Image Source: NeuroscienceNews.com image is credited to S. Seshamani, et al.
Original Research: Abstract for “Detecting default mode networks in utero by integrated 4D fMRI reconstruction and analysis” by Sharmishtaa Seshamani, Anna I. Blazejewska, Susan Mckown, Jason Caucutt, Manjiri Dighe, Christopher Gatenby, and Colin Studholme in Human Brain Mapping. Published online August 11 2016 doi:10.1002/hbm.23303
[cbtabs][cbtab title=”MLA”]NIH/NIBIB. “New Technology Reveals Fetal Brain Activity.” NeuroscienceNews. NeuroscienceNews, 13 October 2016.
<https://neurosciencenews.com/fetal-brain-activity-neuroimaging-5287/>.[/cbtab][cbtab title=”APA”]NIH/NIBIB. (2016, October 13). New Technology Reveals Fetal Brain Activity. NeuroscienceNews. Retrieved October 13, 2016 from https://neurosciencenews.com/fetal-brain-activity-neuroimaging-5287/[/cbtab][cbtab title=”Chicago”]NIH/NIBIB. “New Technology Reveals Fetal Brain Activity.” https://neurosciencenews.com/fetal-brain-activity-neuroimaging-5287/ (accessed October 13, 2016).[/cbtab][/cbtabs]
Abstract
Detecting default mode networks in utero by integrated 4D fMRI reconstruction and analysis
Recently, there has been considerable interest, especially for in utero imaging, in the detection of functional connectivity in subjects whose motion cannot be controlled while in the MRI scanner. These cases require two advances over current studies: (1) multiecho acquisitions and (2) post processing and reconstruction that can deal with significant between slice motion during multislice protocols to allow for the ability to detect temporal correlations introduced by spatial scattering of slices into account. This article focuses on the estimation of a spatially and temporally regular time series from motion scattered slices of multiecho fMRI datasets using a full four-dimensional (4D) iterative image reconstruction framework. The framework which includes quantitative MRI methods for artifact correction is evaluated using adult studies with and without motion to both refine parameter settings and evaluate the analysis pipeline. ICA analysis is then applied to the 4D image reconstruction of both adult and in utero fetal studies where resting state activity is perturbed by motion. Results indicate quantitative improvements in reconstruction quality when compared to the conventional 3D reconstruction approach (using simulated adult data) and demonstrate the ability to detect the default mode network in moving adults and fetuses with single-subject and group analysis.
“Detecting default mode networks in utero by integrated 4D fMRI reconstruction and analysis” by Sharmishtaa Seshamani, Anna I. Blazejewska, Susan Mckown, Jason Caucutt, Manjiri Dighe, Christopher Gatenby, and Colin Studholme in Human Brain Mapping. Published online August 11 2016 doi:10.1002/hbm.23303