Summary: Researchers have created a new map that highlights protein associations through brain networks. The map is available through a software platform that allows users to visualize disease risk factors.
Source: USC.
A study of protein interactions could be the first step to finding treatments that focus on problematic pathways.
Just as parents are not the root of all their children’s problems, a single gene mutation can’t be blamed for complex brain disorders like autism, according to a Keck School of Medicine of USC neuroscientist.
To help researchers see the big picture, Marcelo P. Coba created the first map that highlights the brain’s network of protein associations. It’s a first step to developing treatment drugs that operate more like rifles than shotguns.
“The drugs we have now are not working for these brain disorders,” said Coba, senior author of a new study and an assistant professor of psychiatry at the Zilkha Neurogenetic Institute at the Keck School of Medicine.
“Scientists have not developed a new drug target for complex brain diseases in nearly 60 years. The protein map software my colleagues and I created can help researchers create new therapies that hone in on problem pathways.”
The study was published in late June in Nature Neuroscience. Coba and his colleagues isolated 2,876 protein interactions and figured out where in the brain the protein networks lived, how they communicated and at what age in development those pathways became activated.
Researchers stuffed all that information into a software platform that enables users to visualize disease risk factors throughout the brain’s protein networks.
Taking off the blinders
Many current studies scan patients’ genetics to identify problem genes they label as “risk factors” for developing a disorder.
“The problem is that there is a collection of risk factors contributing to brain disorders,” Coba said. “A single risk factor might explain a very low percentage of the population — perhaps 2 percent of those who have the disease.”
Coba used an analogy. If all flights at a Texas airport were grounded, flight schedules and airports across the country would be affected. A disruption in one location cannot be sustained in that region because the flights are connected in a network of airports, he said.
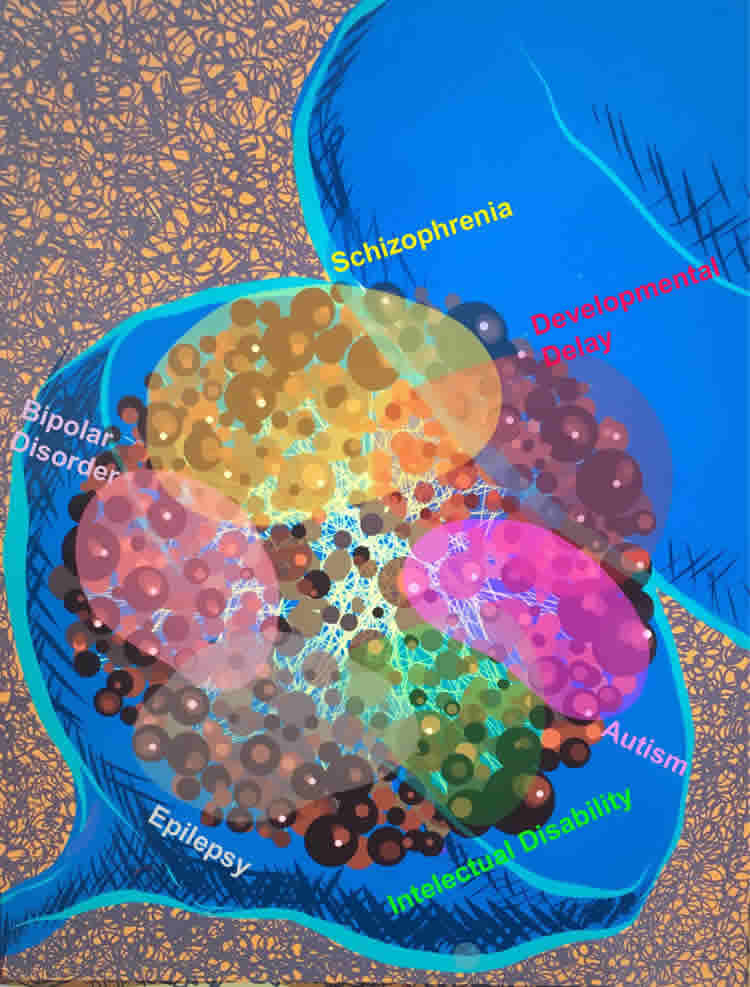
Similarly, genes produce proteins that interact in a protein network. If a gene is mutated, the protein’s connections may experience delays or disruptions. The disorganized protein-to-protein connections from point A to B to C might be the bedrock of brain disorders such as autism, bipolar disorder and schizophrenia, Coba said.
The new software platform is available here.
Funding: The study was supported by $420,000 from the National Institute of Child Health and Human Development (MH104603-01), the National Institutes of Health (MH108728) and the Simons Foundation Autism Research Initiative (248429 and 345034). Seventy percent of research funding originated from the federal government.
Source: Zen Vuong – USC
Image Source: NeuroscienceNews.com image is credited to Steven Park.
Original Research: Abstract for “Spatiotemporal profile of postsynaptic interactomes integrates components of complex brain disorders” by Jing Li, Wangshu Zhang, Hui Yang, Daniel P Howrigan, Brent Wilkinson, Tade Souaiaia, Oleg V Evgrafov, Giulio Genovese, Veronica A Clementel, Jennifer C Tudor, Ted Abel, James A Knowles, Benjamin M Neale, Kai Wang, Fengzhu Sun & Marcelo P Coba in Nature Neuroscience. Published online June 26 2017 doi:10.1038/nn.4594
[cbtabs][cbtab title=”MLA”]USC “New Map May Lead to Drug Development for Complex Brain Disorders.” NeuroscienceNews. NeuroscienceNews, 24 July 2017.
<https://neurosciencenews.com/drug-development-map-7167/>.[/cbtab][cbtab title=”APA”]USC (2017, July 24). New Map May Lead to Drug Development for Complex Brain Disorders. NeuroscienceNew. Retrieved July 24, 2017 from https://neurosciencenews.com/drug-development-map-7167/[/cbtab][cbtab title=”Chicago”]USC “New Map May Lead to Drug Development for Complex Brain Disorders.” https://neurosciencenews.com/drug-development-map-7167/ (accessed July 24, 2017).[/cbtab][/cbtabs]
Abstract
Spatiotemporal profile of postsynaptic interactomes integrates components of complex brain disorders
The postsynaptic density (PSD) contains a collection of scaffold proteins used for assembling synaptic signaling complexes. However, it is not known how the core-scaffold machinery associates in protein-interaction networks or how proteins encoded by genes involved in complex brain disorders are distributed through spatiotemporal protein complexes. Here using immunopurification, proteomics and bioinformatics, we isolated 2,876 proteins across 41 in vivo interactomes and determined their protein domain composition, correlation to gene expression levels and developmental integration to the PSD. We defined clusters for enrichment of schizophrenia, autism spectrum disorders, developmental delay and intellectual disability risk factors at embryonic day 14 and adult PSD in mice. Mutations in highly connected nodes alter protein–protein interactions modulating macromolecular complexes enriched in disease risk candidates. These results were integrated into a software platform, Synaptic Protein/Pathways Resource (SyPPRes), enabling the prioritization of disease risk factors and their placement within synaptic protein interaction networks.
“Spatiotemporal profile of postsynaptic interactomes integrates components of complex brain disorders” by Jing Li, Wangshu Zhang, Hui Yang, Daniel P Howrigan, Brent Wilkinson, Tade Souaiaia, Oleg V Evgrafov, Giulio Genovese, Veronica A Clementel, Jennifer C Tudor, Ted Abel, James A Knowles, Benjamin M Neale, Kai Wang, Fengzhu Sun & Marcelo P Coba in Nature Neuroscience. Published online June 26 2017 doi:10.1038/nn.4594