Summary: Researchers are utilizing neuroimaging data to identify brain patterns and predict depression with the help of machine learning.
Source: University of Texas at Austin.
University of Texas researchers use Stampede supercomputer to identify patterns in neuroimaging data that are predictive for mental disorders.
Depression affects more than 15 million American adults, or about 6.7 percent of the U.S. population, each year. It is the leading cause of disability for those between the ages of 15 and 44.
Is it possible to detect who might be vulnerable to the illness before its onset using brain imaging?
David Schnyer, a cognitive neuroscientist and professor of psychology at The University of Texas at Austin, believes it may be. But identifying its tell-tale signs is no simpler matter. He is using the Stampede supercomputer at the Texas Advanced Computing Center (TACC) to train a machine learning algorithm that can identify commonalities among hundreds of patients using Magnetic Resonance Imaging (MRI) brain scans, genomics data and other relevant factors, to provide accurate predictions of risk for those with depression and anxiety.
Researchers have long studied mental disorders by examining the relationship between brain function and structure in neuroimaging data.
“One difficulty with that work is that it’s primarily descriptive. The brain networks may appear to differ between two groups, but it doesn’t tell us about what patterns actually predict which group you will fall into,” Schnyer says. “We’re looking for diagnostic measures that are predictive for outcomes like vulnerability to depression or dementia.”
In 2017, Schnyer, working with Peter Clasen (University of Washington School of Medicine), Christopher Gonzalez (University of California, San Diego) and Christopher Beevers (UT Austin), completed their analysis of a proof-of-concept study that used a machine learning approach to classify individuals with major depressive disorder with roughly 75 percent accuracy.
Machine learning is a subfield of computer science that involves the construction of algorithms that can “learn” by building a model from sample data inputs, and then make independent predictions on new data.
The type of machine learning that Schnyer and his team tested is called Support Vector Machine Learning. The researchers provided a set of training examples, each marked as belonging to either healthy individuals or those who have been diagnosed with depression. Schnyer and his team labelled features in their data that were meaningful, and these examples were used to train the system. A computer then scanned the data, found subtle connections between disparate parts, and built a model that assigns new examples to one category or the other.
In the study, Schnyer analyzed brain data from 52 treatment-seeking participants with depression, and 45 heathy control participants. To compare the groups, they matched a subset of depressed participants with healthy individuals based on age and gender, bringing the sample size to 50.
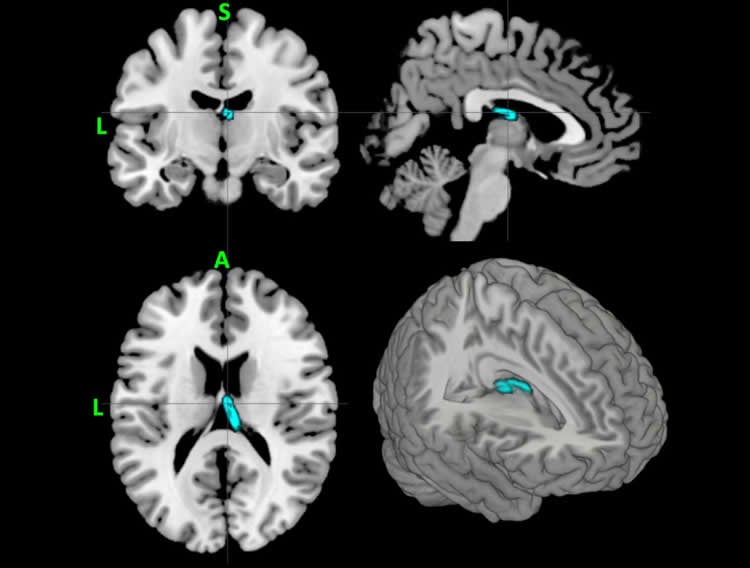
Participants received diffusion tensor imaging (DTI) MRI scans, which tag water molecules to determine the extent to which those molecules are microscopically diffused in the brain over time. By measuring this diffusion in multiple spatial directions, vectors are generated for each voxel (three-dimensional cubes that represent either structure or neural activity throughout the brain) to quantify the dominant fiber orientation. These measurements are then translated into metrics that indicate the integrity of white matter pathways within the cerebral cortex.
One common parameter used to characterize DTI is fractional anisotropy: the extent to which diffusion is highly directional (high fractional anisotropy) or unrestricted (low fractional anisotropy).
They compared these fractional anisotropy measurements between the two groups and found statistically significant differences. They then reduced the number of voxels involved to a subset that was most relevant for classification and carried out the classification and prediction using the machine learning approach.
“We feed in whole brain data or a subset and predict disease classifications or any potential behavioral measure such as measures of negative information bias,” he says.
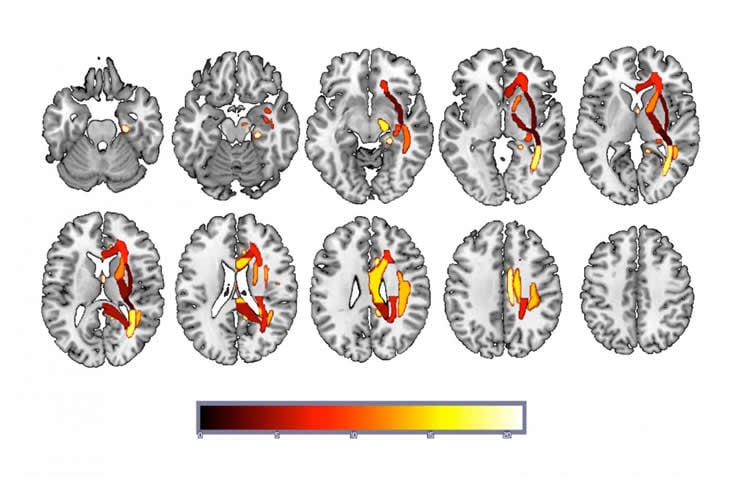
The study revealed that DTI-derived fractional anisotropy maps can accurately classify depressed or vulnerable individuals versus healthy controls. It also showed that predictive information is distributed across brain networks rather than being highly localized.
“Not only are were learning that we can classify depressed versus non-depressed people using DTI data, we are also learning something about how depression is represented within the brain,” said Beevers, a professor of psychology and director of the Institute for Mental Health Research at UT Austin. “Rather than trying to find the area that is disrupted in depression, we are learning that alterations across a number of networks contribute to the classification of depression.”
The scale and complexity of the problem necessitates a machine learning approach. Each brain is represented by roughly 175,000 voxels and detecting complex relationship among such a large number of components by looking at the scans is practically impossible. For that reason, the team uses machine learning to automate the discovery process.
“This is the wave of the future,” Schnyer says. “We’re seeing increasing numbers of articles and presentations at conference on the application of machine learning to solve difficult problems in neuroscience.”
The results are promising, but not yet clear-cut enough to be used as a clinical metric. However, Schnyer believes that by adding more data — related not only to MRI scans but also from genomics and other classifiers — the system can do much better.
“One of the benefits of machine learning, compared to more traditional approaches, is that machine learning should increase the likelihood that what we observe in our study will apply to new and independent datasets. That is, it should generalize to new data,” Beevers said. “This is a critical question that we are really excited to test in future studies.”
Beevers and Schnyer will expand their study to include data from several hundred volunteers from the Austin community who have been diagnosed with depression, anxiety or a related condition. Stampede 2 — TACC’s newest supercomputer which will come online later in 2017 and will be twice as powerful as the current system — will provide the increased computer processing power required to incorporate more data and achieve greater accuracy.
“This approach, and also the movement towards open science and large databases like the human connectome project, mean that facilities like TACC are absolutely essential,” Schnyer says. “You just can’t do this work on desktops. It’s going to become more and more important to have an established relationship with an advanced computing center.”
Funding: National Institutes of Health, National Science Foundation funded this study.
Source: Aaron Dubrow – University of Texas at Austin
Image Source: NeuroscienceNews.com images are credited to David M Schnyer, Peter C. Clasen, Christopher Gonzalez and Christopher G. Beevers.
Original Research: Abstract for “Evaluating the diagnostic utility of applying a machine learning algorithm to diffusion tensor MRI measures in individuals with major depressive disorder” by David M Schnyer, Peter C. Clasen, Christopher Gonzalez, and Christopher G. Beevers in Psychiatry Research: Neuroimaging. Published online March 22 2017 doi:10.1016/j.pscychresns.2017.03.003
[cbtabs][cbtab title=”MLA”]University of Texas at Austin “Enlisting The Help of Machine Learning to Diagnose Depression.” NeuroscienceNews. NeuroscienceNews, 28 March 2017.
<https://neurosciencenews.com/depression-machine-learning-6301/>.[/cbtab][cbtab title=”APA”]University of Texas at Austin (2017, March 28). Enlisting The Help of Machine Learning to Diagnose Depression. NeuroscienceNew. Retrieved March 28, 2017 from https://neurosciencenews.com/depression-machine-learning-6301/[/cbtab][cbtab title=”Chicago”]University of Texas at Austin “Enlisting The Help of Machine Learning to Diagnose Depression.” https://neurosciencenews.com/depression-machine-learning-6301/ (accessed March 28, 2017).[/cbtab][/cbtabs]
Abstract
Evaluating the diagnostic utility of applying a machine learning algorithm to diffusion tensor MRI measures in individuals with major depressive disorder
Highlights
•Examination of utility of applying support vector machine (SVM) learning to MRI measures of brain white matter in order to classify individuals with major depressive disorder (MDD).
•High degree of accuracy when utilizing a feature selection approach.
•Distributed information is more diagnostically informative than highly localized.
Abstract
Using MRI to diagnose mental disorders has been a long-term goal. Despite this, the vast majority of prior neuroimaging work has been descriptive rather than predictive. The current study applies support vector machine (SVM) learning to MRI measures of brain white matter to classify adults with Major Depressive Disorder (MDD) and healthy controls. In a precisely matched group of individuals with MDD (n = 25) and healthy controls (n = 25), SVM learning accurately (74%) classified patients and controls across a brain map of white matter fractional anisotropy values (FA). The study revealed three main findings: 1) SVM applied to DTI derived FA maps can accurately classify MDD vs. healthy controls; 2) prediction is strongest when only right hemisphere white matter is examined; and 3) removing FA values from a region identified by univariate contrast as significantly different between MDD and healthy controls does not change the SVM accuracy. These results indicate that SVM learning applied to neuroimaging data can classify the presence versus absence of MDD and that predictive information is distributed across brain networks rather than being highly localized. Finally, MDD group differences revealed through typical univariate contrasts do not necessarily reveal patterns that provide accurate predictive information.
“Evaluating the diagnostic utility of applying a machine learning algorithm to diffusion tensor MRI measures in individuals with major depressive disorder” by David M Schnyer, Peter C. Clasen, Christopher Gonzalez, and Christopher G. Beevers in Psychiatry Research: Neuroimaging. Published online March 22 2017 doi:10.1016/j.pscychresns.2017.03.003