Summary: SARS-CoV-2 is detectable in aerosols for up to three hours and up to four hours on copper. On cardboard, the virus is detectable for up to 24 hours, and up to three days on plastic and stainless steel. The study suggests the virus may be acquired through the air in addition to touching surfaces.
Source: NIH/NIAID
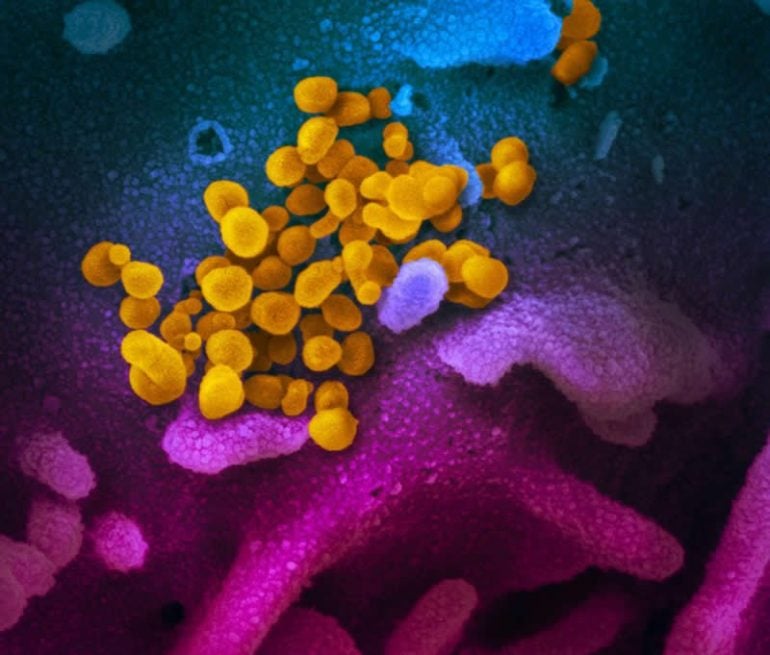
The virus that causes coronavirus disease 2019 (COVID-19) is stable for several hours to days in aerosols and on surfaces, according to a new study from National Institutes of Health, CDC, UCLA and Princeton University scientists The New England Journal of Medicine. The scientists found that severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) was detectable in aerosols for up to three hours, up to four hours on copper, up to 24 hours on cardboard and up to two to three days on plastic and stainless steel. The results provide key information about the stability of SARS-CoV-2, which causes COVID-19 disease, and suggests that people may acquire the virus through the air and after touching contaminated objects. The study information was widely shared during the past two weeks after the researchers placed the contents on a preprint server to quickly share their data with colleagues.
The NIH scientists, from the National Institute of Allergy and Infectious Diseases’ Montana facility at Rocky Mountain Laboratories, compared how the environment affects SARS-CoV-2 and SARS-CoV-1, which causes SARS. SARS-CoV-1, like its successor now circulating across the globe, emerged from China and infected more than 8,000 people in 2002 and 2003. SARS-CoV-1 was eradicated by intensive contact tracing and case isolation measures and no cases have been detected since 2004. SARS-CoV-1 is the human coronavirus most closely related to SARS-CoV-2. In the stability study the two viruses behaved similarly, which unfortunately fails to explain why COVID-19 has become a much larger outbreak.
The NIH study attempted to mimic virus being deposited from an infected person onto everyday surfaces in a household or hospital setting, such as through coughing or touching objects. The scientists then investigated how long the virus remained infectious on these surfaces.

The scientists highlighted additional observations from their study:
- If the viability of the two coronaviruses is similar, why is SARS-CoV-2 resulting in more cases? Emerging evidence suggests that people infected with SARS-CoV-2 might be spreading virus without recognizing, or prior to recognizing, symptoms. This would make disease control measures that were effective against SARS-CoV-1 less effective against its successor.
- In contrast to SARS-CoV-1, most secondary cases of virus transmission of SARS-CoV-2 appear to be occurring in community settings rather than healthcare settings. However, healthcare settings are also vulnerable to the introduction and spread of SARS-CoV-2, and the stability of SARS-CoV-2 in aerosols and on surfaces likely contributes to transmission of the virus in healthcare settings.
The findings affirm the guidance from public health professionals to use precautions similar to those for influenza and other respiratory viruses to prevent the spread of SARS-CoV-2:
- Avoid close contact with people who are sick.
- Avoid touching your eyes, nose, and mouth.
- Stay home when you are sick.
- Cover your cough or sneeze with a tissue, then throw the tissue in the trash.
- Clean and disinfect frequently touched objects and surfaces using a regular household cleaning spray or wipe.
Source:
NIH/NIAID
Media Contacts:
Ken Pekoc – NIH/NIAID
Image Source:
The image is credited to NIAID RML.
Original Research: Open access
“Aerosol and surface stability of HCoV-19 (SARS-CoV-2) compared to SARS-CoV-1”. N. van Doremalen, et al.
New England Journal of Medicine doi:10.1056/NEJMc2004973.
Abstract
Aerosol and surface stability of HCoV-19 (SARS-CoV-2) compared to SARS-CoV-1
A novel human coronavirus that is now named severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) (formerly called HCoV-19) emerged in Wuhan, China, in late 2019 and is now causing a pandemic. We analyzed the aerosol and surface stability of SARS-CoV-2 and compared it with SARS-CoV-1, the most closely related human coronavirus.
We evaluated the stability of SARS-CoV-2 and SARS-CoV-1 in aerosols and on various surfaces and estimated their decay rates using a Bayesian regression model (see the Methods section in the Supplementary Appendix, available with the full text of this letter at NEJM.org). SARS-CoV-2 nCoV-WA1-2020 (MN985325.1) and SARS-CoV-1 Tor2 (AY274119.3) were the strains used. Aerosols (<5 μm) containing SARS-CoV-2 (105.25 50% tissue-culture infectious dose [TCID50] per milliliter) or SARS-CoV-1 (106.75-7.00 TCID50 per milliliter) were generated with the use of a three-jet Collison nebulizer and fed into a Goldberg drum to create an aerosolized environment. The inoculum resulted in cycle-threshold values between 20 and 22, similar to those observed in samples obtained from the upper and lower respiratory tract in humans.