The goal of brain simulations using supercomputers is to understand the processes in our brain. This is a mammoth task: the activity of an estimated 100 billion nerve cells – also known as neurons – must be represented . It is also a task that has historically been impossible because even the most powerful computers in the world can only simulate one percent of the nerve cells due to memory constraints. For this reason, scientists have turned to downscaled models. However, this downscaling is problematic, as shown by a recent Juelich study published in PLOS Computational Biology.
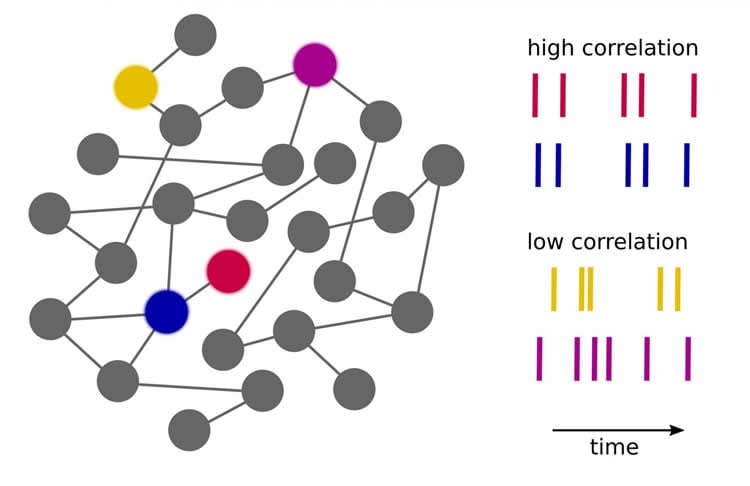
“The challenge of brain simulation is that the nerve cells enter into a temporary relationship with other neurons depending on the task at hand,” says Prof. Dr. Markus Diesmann, director of Juelich’s institute Computational and Systems Neuroscience (INM-6). Every nerve cell is linked to an average of 10,000 other neurons, which synchronize their activity with each other to varying degrees. The intensity of the relationship between neurons – referred to as correlation – varies depending on the task and the brain areas involved. Using mathematical methods, Dr. Sacha van Albada, research scientist in Markus Diesmann’s lab, Dr. Moritz Helias, head of the group Theory of Multi-Scale Neuronal Networks, and Diesmann demonstrated that this correlation cannot be correctly preserved when the number of neuronal connections in a brain model is below a certain level. However, correlations form the basis of frequently used, measurable signals in the brain such as the EEG and the local field potential (LFP).
Each neuron has around 10,000 connections
The information flow in the human brain is extremely complex. Nerve cells exchange information with each other in the form of electrical signals via so-called synapses. Each nerve cell has around 10,000 such connections through which it communicates with other neurons. Just as the motorway network does not determine which car should drive where, data in the brain choose different routes and exits depending on the task. Today’s computers cannot process or store such gigantic quantities of information. The number of synapses is therefore reduced in many brain models, which in turn diminishes memory usage.
The Human Brain Project (HBP) aims at detailed simulations of the human brain
Despite these difficulties, the detailed simulation of the entire human brain on a future supercomputer remains the aim of a large-scale scientific project. In the EU-funded Human Brain Project (HBP), neuroscientists and physicists like Markus Diesmann are working together with computer scientists, medical scientists, and mathematicians from over 80 European and international scientific institutions. “Our current research work is a further indication that there is no way around simulating brain circuits in their natural size if we want to gain solid knowledge,” says Diesmann.
One of the biggest challenges of the Human Brain Project is the development of new supercomputers. Scientists from Juelich also have a leading role here: the Juelich Supercomputing Centre (JSC) is developing exascale computers to perform the complex simulations in the Human Brain Project. This requires a 100-fold increase in the computing power of today’s supercomputers. Next to the mathematical modeling as in the recent study Markus Diesmann and his team therefore work in parallel on the creation of simulation software for the new generation of computers. This work is being performed at the institute Theoretical Neuroscience (IAS-6) and as part of the Neural Simulation Technology Initiative, which provides free access to the software NEST.
Source: Tobias Schloesser – Forschungszentrum Jülich
Image Source: The image is credited to Forschungszentrum Jülich
Original Research: Full open access research for “Scalability of Asynchronous Networks Is Limited by One-to-One Mapping between Effective Connectivity and Correlations” by Sacha Jennifer van Albada, Moritz Helias, and Markus Diesmann in PLOS Computational Biology. Published online September 1 2015 doi:10.1371/journal.pcbi.1004490
Abstract
Scalability of Asynchronous Networks Is Limited by One-to-One Mapping between Effective Connectivity and Correlations
Network models are routinely downscaled compared to nature in terms of numbers of nodes or edges because of a lack of computational resources, often without explicit mention of the limitations this entails. While reliable methods have long existed to adjust parameters such that the first-order statistics of network dynamics are conserved, here we show that limitations already arise if also second-order statistics are to be maintained. The temporal structure of pairwise averaged correlations in the activity of recurrent networks is determined by the effective population-level connectivity. We first show that in general the converse is also true and explicitly mention degenerate cases when this one-to-one relationship does not hold. The one-to-one correspondence between effective connectivity and the temporal structure of pairwise averaged correlations implies that network scalings should preserve the effective connectivity if pairwise averaged correlations are to be held constant. Changes in effective connectivity can even push a network from a linearly stable to an unstable, oscillatory regime and vice versa. On this basis, we derive conditions for the preservation of both mean population-averaged activities and pairwise averaged correlations under a change in numbers of neurons or synapses in the asynchronous regime typical of cortical networks. We find that mean activities and correlation structure can be maintained by an appropriate scaling of the synaptic weights, but only over a range of numbers of synapses that is limited by the variance of external inputs to the network. Our results therefore show that the reducibility of asynchronous networks is fundamentally limited.
“Scalability of Asynchronous Networks Is Limited by One-to-One Mapping between Effective Connectivity and Correlations” by Sacha Jennifer van Albada, Moritz Helias, and Markus Diesmann in PLOS Computational Biology doi:10.1371/journal.pcbi.1004490