Breakthrough could help see pathways that degenerate with Parkinson’s and Huntington’s disease.
Certain diseases, like Parkinson’s and Huntington’s disease, are associated with damage to the pathways between the brain’s basal ganglia regions. The basal ganglia sits at the base of the brain and is responsible for, among other things, coordinating movement. It is made up of four interconnected, deep brain structures that imaging techniques have previously been unable to visualize.
For the first time, Carnegie Mellon University BrainHub scientists have used a non-invasive brain-imaging tool to detect the pathways that connect the parts of the basal ganglia. Published in NeuroImage, the research provides a better understanding of this area’s circuitry, which could potentially lead to technologies to help track disease progression for Parkinson’s and Huntington’s disease and other neurological disorders.
“Clinically, it is difficult to see the pathways within the basal ganglia with neuroimaging techniques, like the ever popular MRI, because many of the fiber bundles that make up key parts of this circuit are very small and buried within dense cell bodies,” said Patrick Beukema, the lead author and a graduate student in the Center for Neuroscience at the University of Pittsburgh (CNUP) and the joint Pitt and CMU Center for the Basis of Neural Cognition (CNBC).
“For reasons that are not fully understood, the pathways that connect the basal ganglia’s regions are highly susceptible to damage. Because they are important for motor control, this damage can result in substantial motor deficits, so it is highly desirable to discover more about this area,” Beukema said.
Diffusion MRI measures the movement of water molecules to create a visual representation of the brain’s axons. In this study, the research team used two types of diffusion imaging to visualize the major pathways that connect the internal circuitry of the basal ganglia. Sixty healthy adults had their brains scanned using diffusion spectrum imaging, which provided a picture of the orientation of moving water molecules. And, multi-shell imaging was used on 78 healthy adults to get similar images using different imaging parameters.
The results from both imaging techniques showed that it is possible to detect the small but important fiber connections in the brain. The researchers also found that by looking at the general patterns of water movement in the basal ganglia, they could automatically distinguish one small brain region from the other.
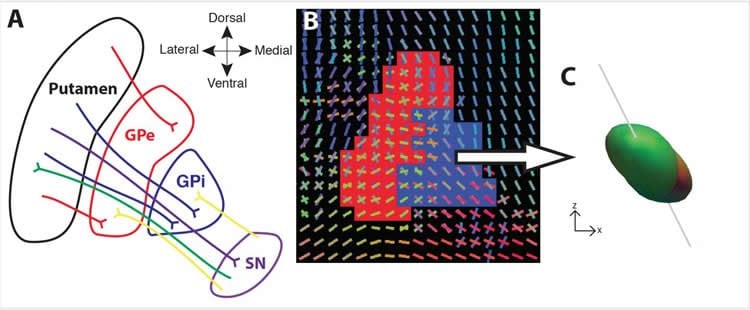
“The pathways that Patrick has been able to visualize are critical to so many functions, yet we haven’t been able to see them in the living human brain before. This opens the door to so many research and clinical opportunities,” said Timothy J. Verstynen, assistant professor of psychology in CMU’s Dietrich College of Humanities and Social Sciences and CNBC faculty member.
This is not the first brain research breakthrough to happen at Carnegie Mellon. CMU is the birthplace of artificial intelligence and cognitive psychology and has been a leader in the study of brain and behavior for more than 50 years. The university has created some of the first cognitive tutors, helped to develop the Jeopardy-winning Watson, founded a groundbreaking doctoral program in neural computation, and completed cutting-edge work in understanding the genetics of autism. Building on its strengths in biology, computer science, psychology, statistics and engineering, CMU launched BrainHub, an initiative that focuses on how the structure and activity of the brain give rise to complex behaviors.
In addition to Beukema and Verstynen, the research team included Fang-Cheng Yeh, a research psychologist in CMU’s Department of Psychology and the CNBC.
Funding: This research was funded by a National Science Foundation BIG DATA grant and support from the Army Research Laboratory and the CNUP.
Source: Shilo Rea – Carnegie Mellon University
Image Source: The image is credited to Carnegie Mellon University
Original Research: Abstract for “In vivo characterization of the connectivity and subcomponents of the human globus pallidus” by Patrick Beukema, Fang-Cheng Yeh, and Timothy Verstynen in Neuroimage: Clinical. Published online July 18 2015 doi:10.1016/j.neuroimage.2015.07.031
Abstract
In vivo characterization of the connectivity and subcomponents of the human globus pallidus
Projections from the substantia nigra and striatum traverse through the pallidum on the way to their targets. To date, in vivo characterization of these pathways remains elusive. Here we used high angular resolution diffusion imaging (N = 138) to study the characteristics and structural subcompartments of the human pallidum. Our central result shows that the diffusion orientation distribution functions within the pallidum are asymmetrically oriented in a dorsal to dorsolateral direction, consistent with the orientation of underlying fiber systems. We also observed systematic differences in the diffusion signal between the two pallidal segments. Compared to the outer pallidal segment, the internal segment has more peaks in the diffusion orientation distribution and stronger anisotropy in the primary fiber direction, consistent with known cellular differences between the underlying nuclei. These differences in orientation, complexity, and degree of anisotropy are sufficiently robust to automatically segment the pallidal nuclei using diffusion properties. We characterize these patterns in one data set using diffusion spectrum imaging and replicate in a separate sample of subjects imaged using multi-shell imaging, highlighting the reliability of these diffusion patterns within pallidal nuclei. Thus the gray matter diffusion signal can be useful as an in vivo measure of the collective efferent pathways running through the human pallidum.
“In vivo characterization of the connectivity and subcomponents of the human globus pallidus” by Patrick Beukema, Fang-Cheng Yeh, and Timothy Verstynen in Neuroimage: Clinical. Published online July 18 2015 doi:10.1016/j.neuroimage.2015.07.031